Summary information and primary citation
- PDB-id
- 7lw8; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- hydrolase-DNA
- Method
- X-ray (2.88 Å)
- Summary
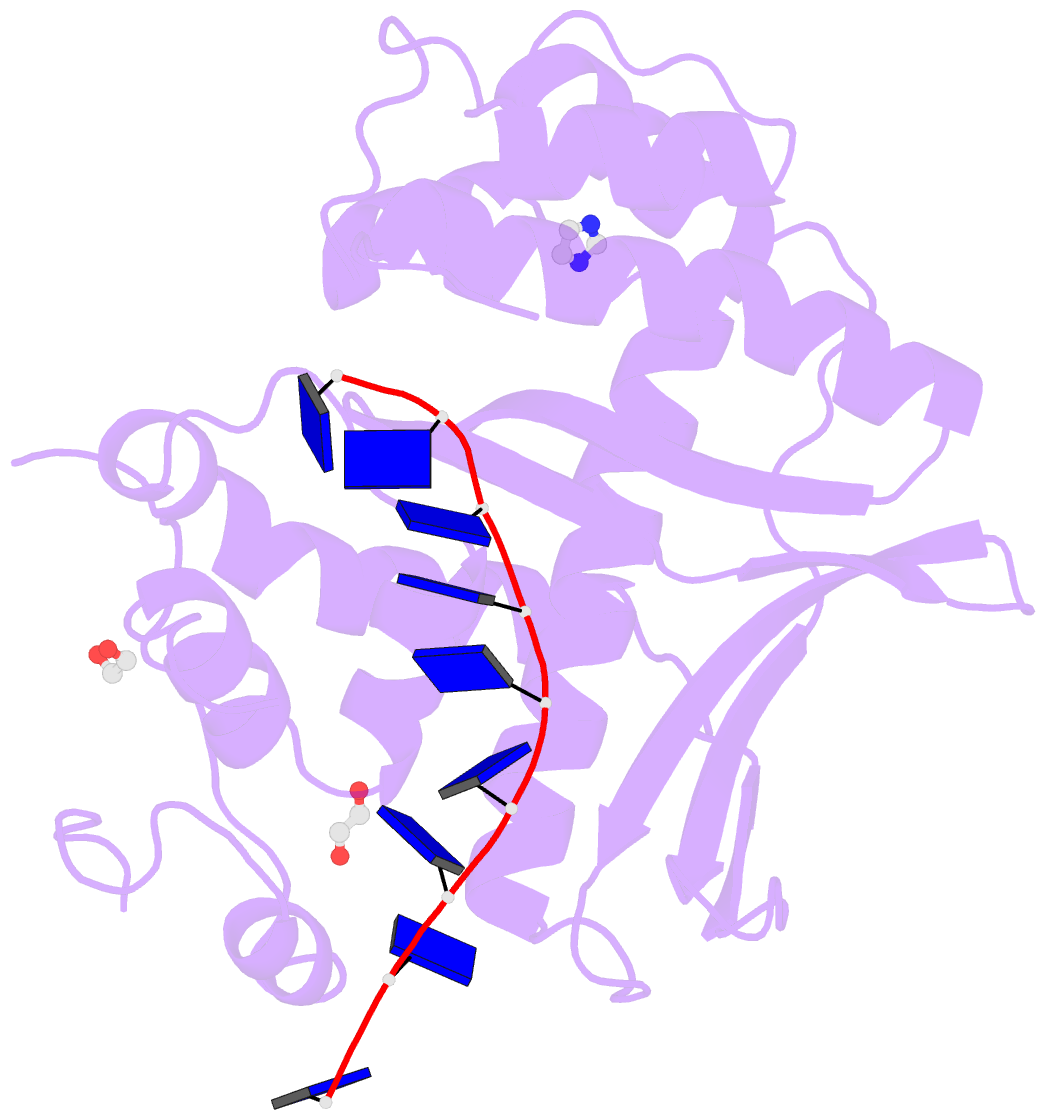
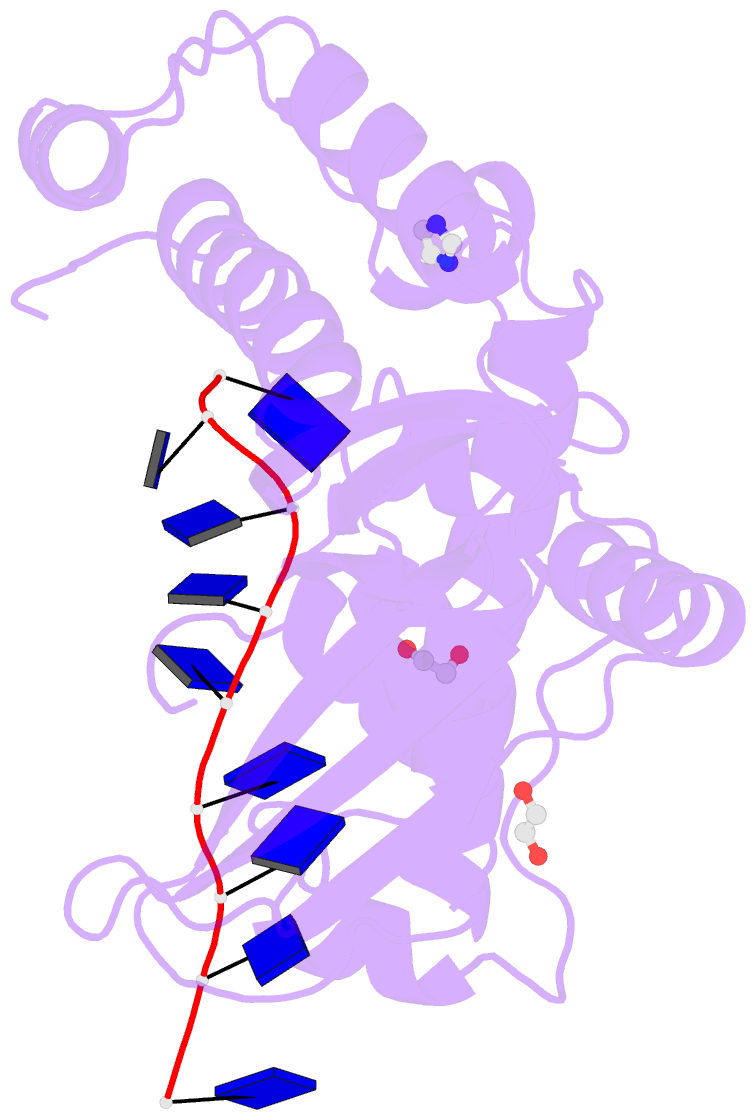
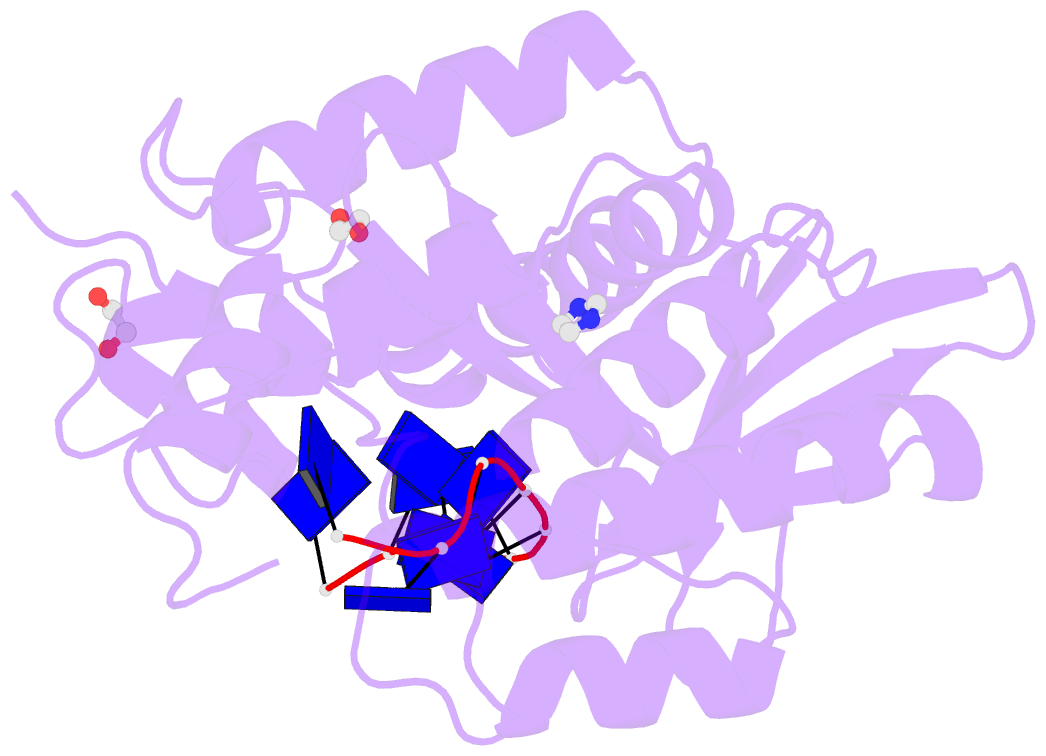
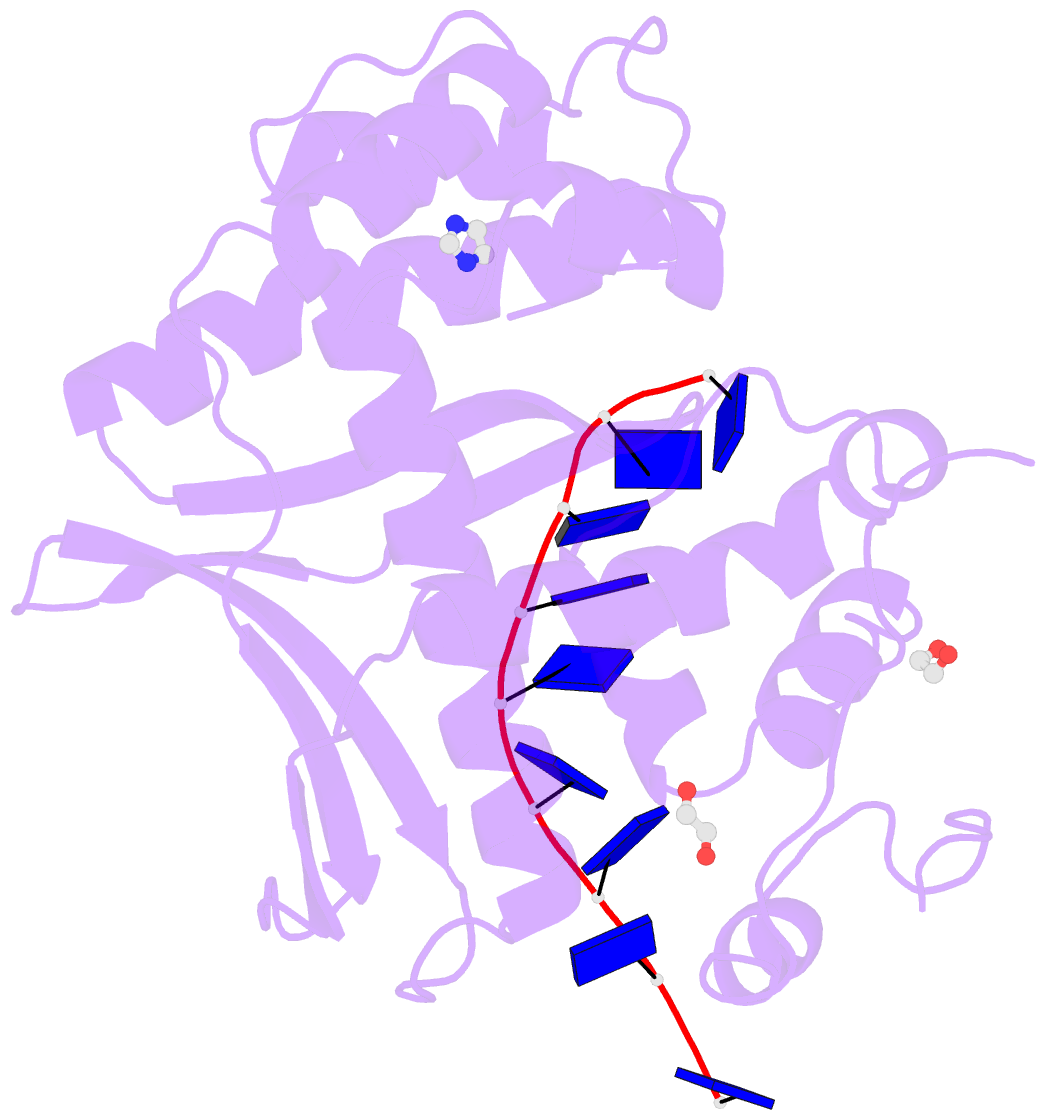
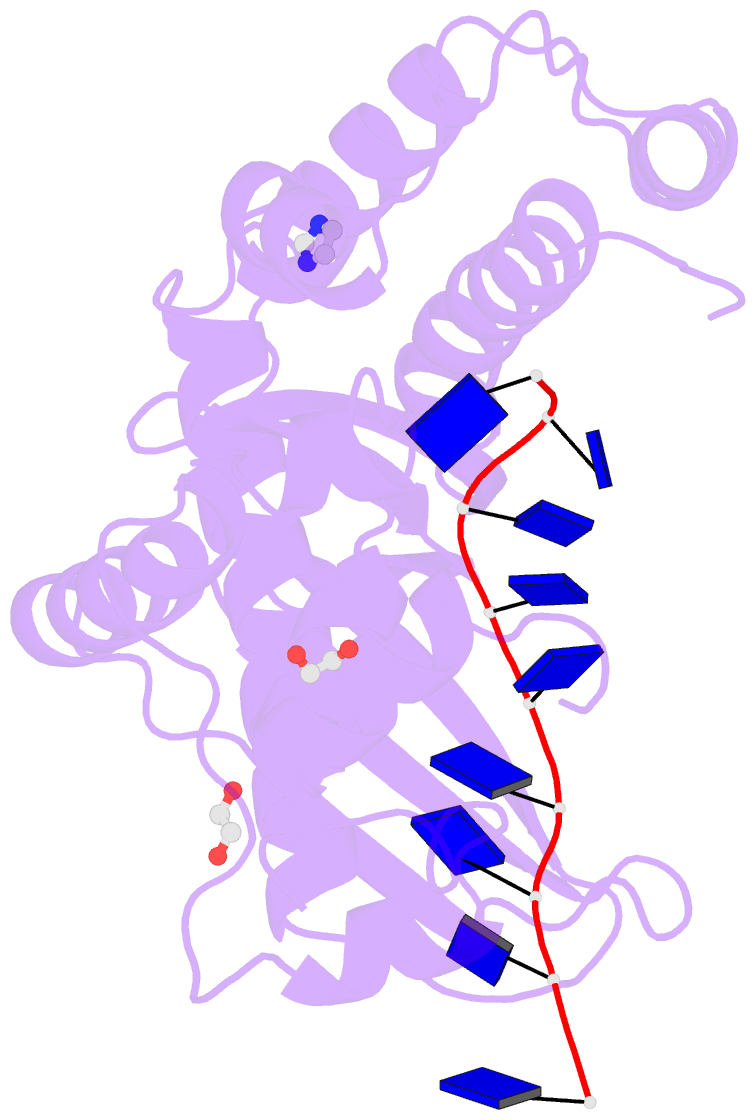
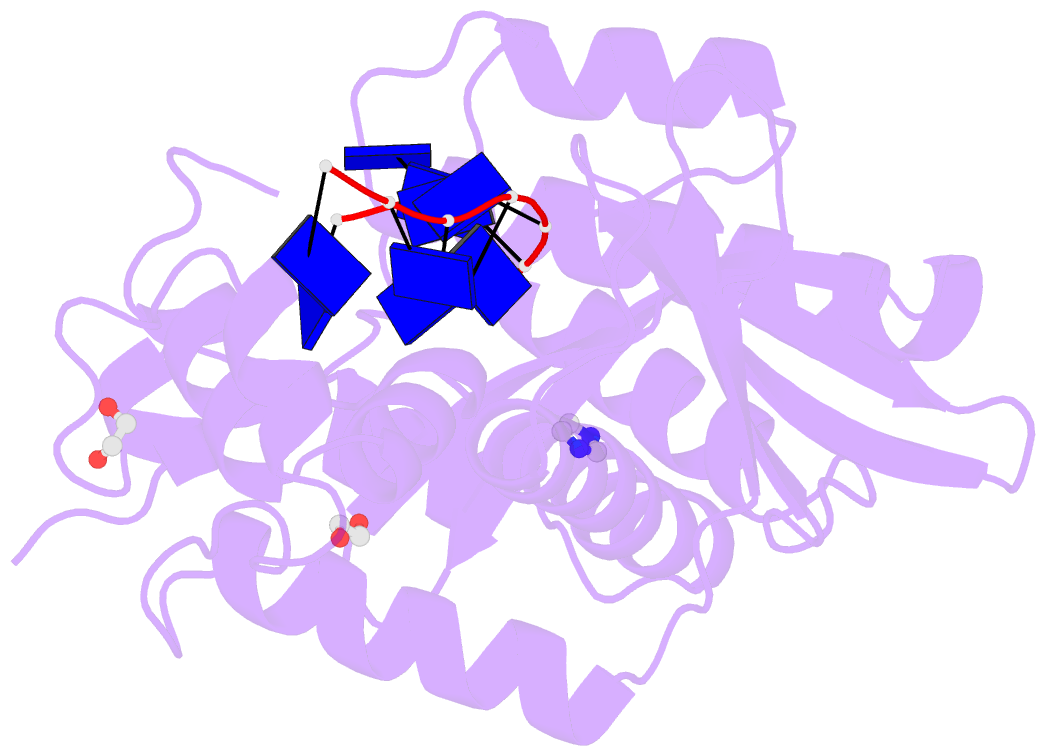
- Human exonuclease 5 crystal structure in complex with a ssDNA
- Reference
- Hambarde S, Tsai CL, Pandita RK, Bacolla A, Maitra A, Charaka V, Hunt CR, Kumar R, Limbo O, Le Meur R, Chazin WJ, Tsutakawa SE, Russell P, Schlacher K, Pandita TK, Tainer JA (2021): "EXO5-DNA structure and BLM interactions direct DNA resection critical for ATR-dependent replication restart." Mol.Cell, 81, 2989. doi: 10.1016/j.molcel.2021.05.027.
- Abstract
- Stalled DNA replication fork restart after stress as orchestrated by ATR kinase, BLM helicase, and structure-specific nucleases enables replication, cell survival, and genome stability. Here we unveil human exonuclease V (EXO5) as an ATR-regulated DNA structure-specific nuclease and BLM partner for replication fork restart. We find that elevated EXO5 in tumors correlates with increased mutation loads and poor patient survival, suggesting that EXO5 upregulation has oncogenic potential. Structural, mechanistic, and mutational analyses of EXO5 and EXO5-DNA complexes reveal a single-stranded DNA binding channel with an adjacent ATR phosphorylation motif (T88Q89) that regulates EXO5 nuclease activity and BLM binding identified by mass spectrometric analysis. EXO5 phospho-mimetic mutant rescues the restart defect from EXO5 depletion that decreases fork progression, DNA damage repair, and cell survival. EXO5 depletion furthermore rescues survival of FANCA-deficient cells and indicates EXO5 functions epistatically with SMARCAL1 and BLM. Thus, an EXO5 axis connects ATR and BLM in directing replication fork restart.