Summary information and primary citation
- PDB-id
- 7qdd; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- RNA binding protein
- Method
- NMR
- Summary
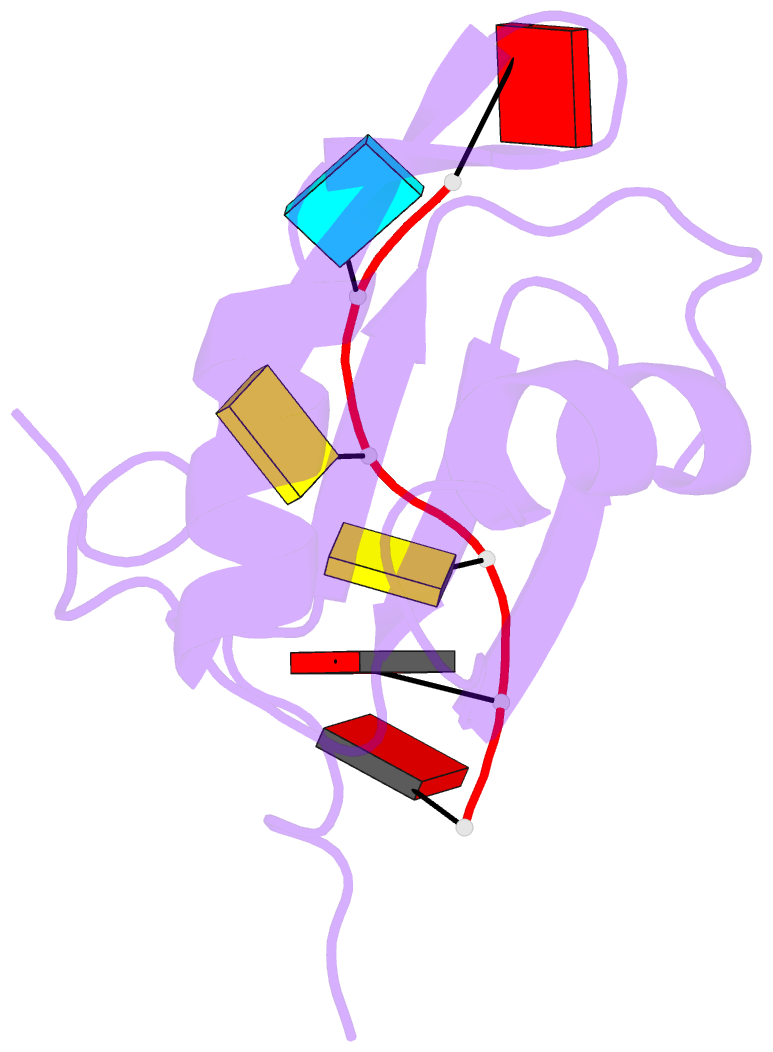
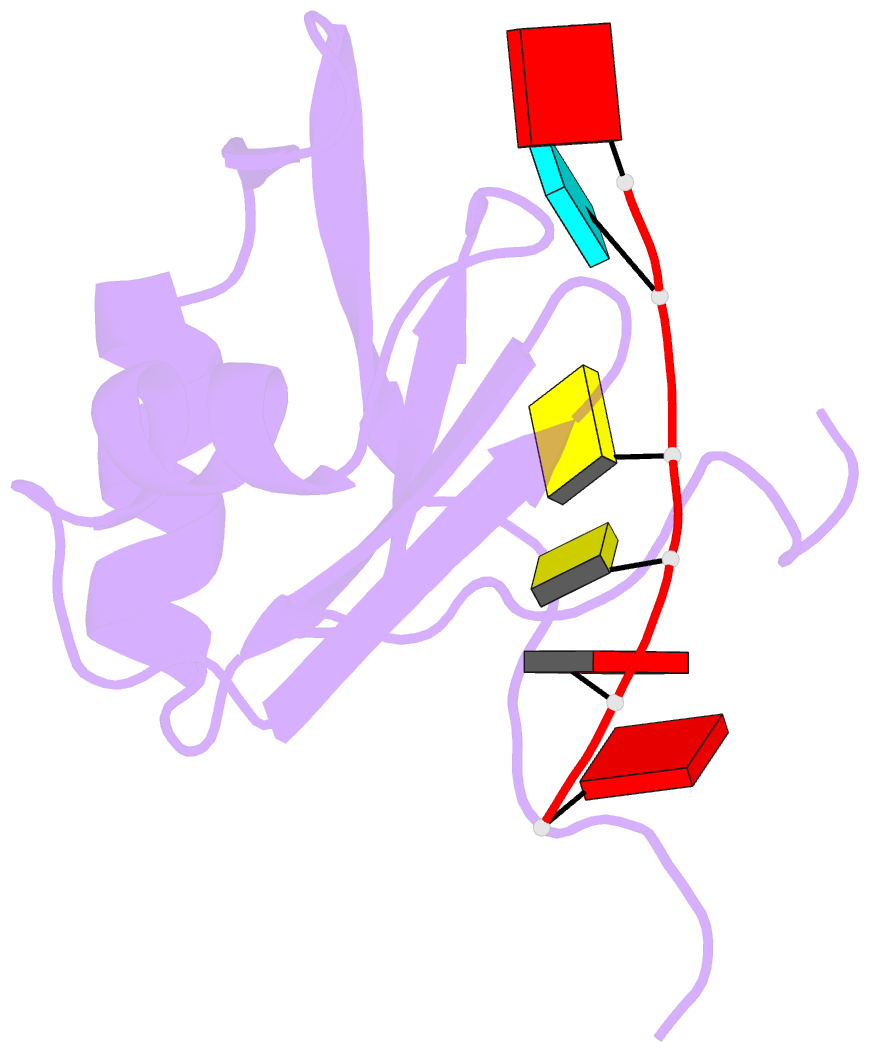
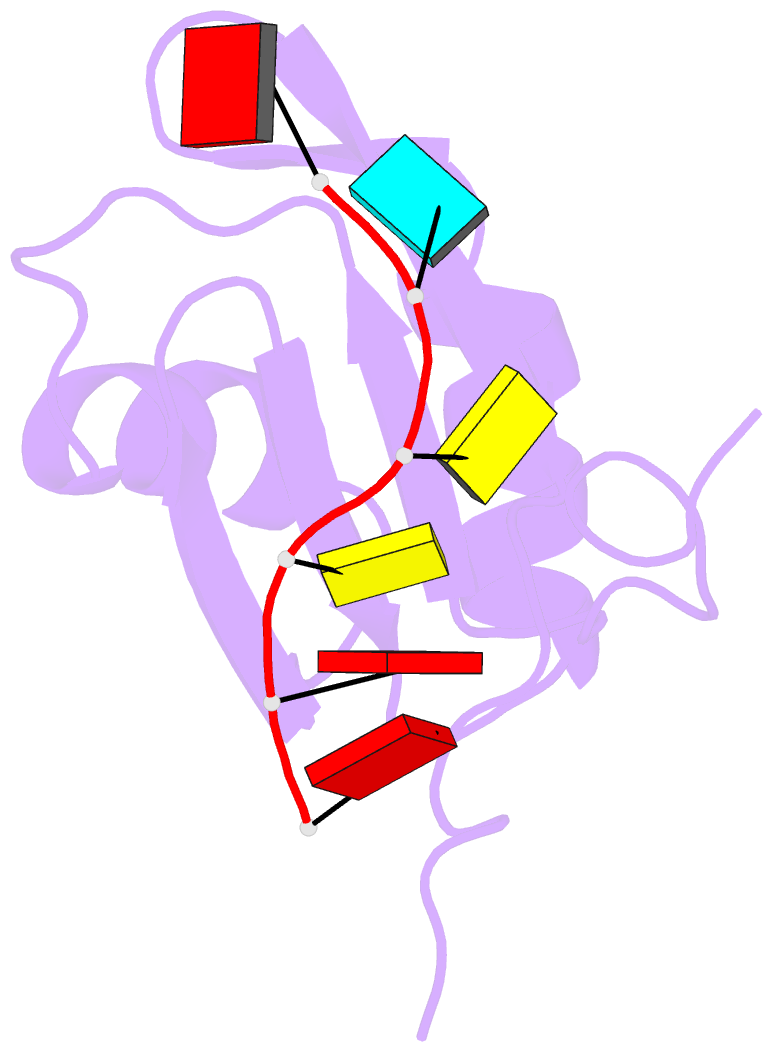
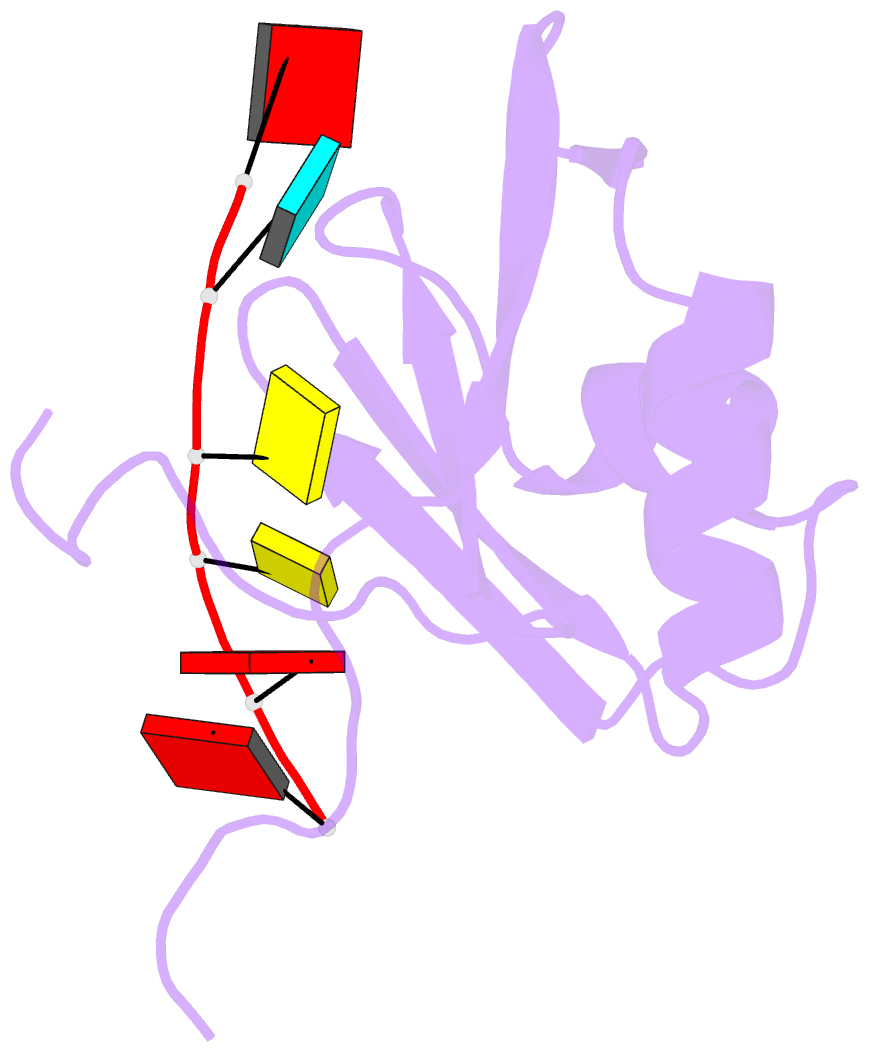
- NMR structure of npl3 rrm1 bound to the auccaa RNA
- Reference
- Moursy A, Clery A, Gerhardy S, Betz KM, Rao S, Mazur J, Campagne S, Beusch I, Duszczyk MM, Robinson MD, Panse VG, Allain FH (2023): "RNA recognition by Npl3p reveals U2 snRNA-binding compatible with a chaperone role during splicing." Nat Commun, 14, 7166. doi: 10.1038/s41467-023-42962-4.
- Abstract
- The conserved SR-like protein Npl3 promotes splicing of diverse pre-mRNAs. However, the RNA sequence(s) recognized by the RNA Recognition Motifs (RRM1 & RRM2) of Npl3 during the splicing reaction remain elusive. Here, we developed a split-iCRAC approach in yeast to uncover the consensus sequence bound to each RRM. High-resolution NMR structures show that RRM2 recognizes a 5´-GNGG-3´ motif leading to an unusual mille-feuille topology. These structures also reveal how RRM1 preferentially interacts with a CC-dinucleotide upstream of this motif, and how the inter-RRM linker and the region C-terminal to RRM2 contribute to cooperative RNA-binding. Structure-guided functional studies show that Npl3 genetically interacts with U2 snRNP specific factors and we provide evidence that Npl3 melts U2 snRNA stem-loop I, a prerequisite for U2/U6 duplex formation within the catalytic center of the Bact spliceosomal complex. Thus, our findings suggest an unanticipated RNA chaperoning role for Npl3 during spliceosome active site formation.