Summary information and primary citation
- PDB-id
- 7xc7; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- RNA binding protein-RNA
- Method
- cryo-EM (3.1 Å)
- Summary
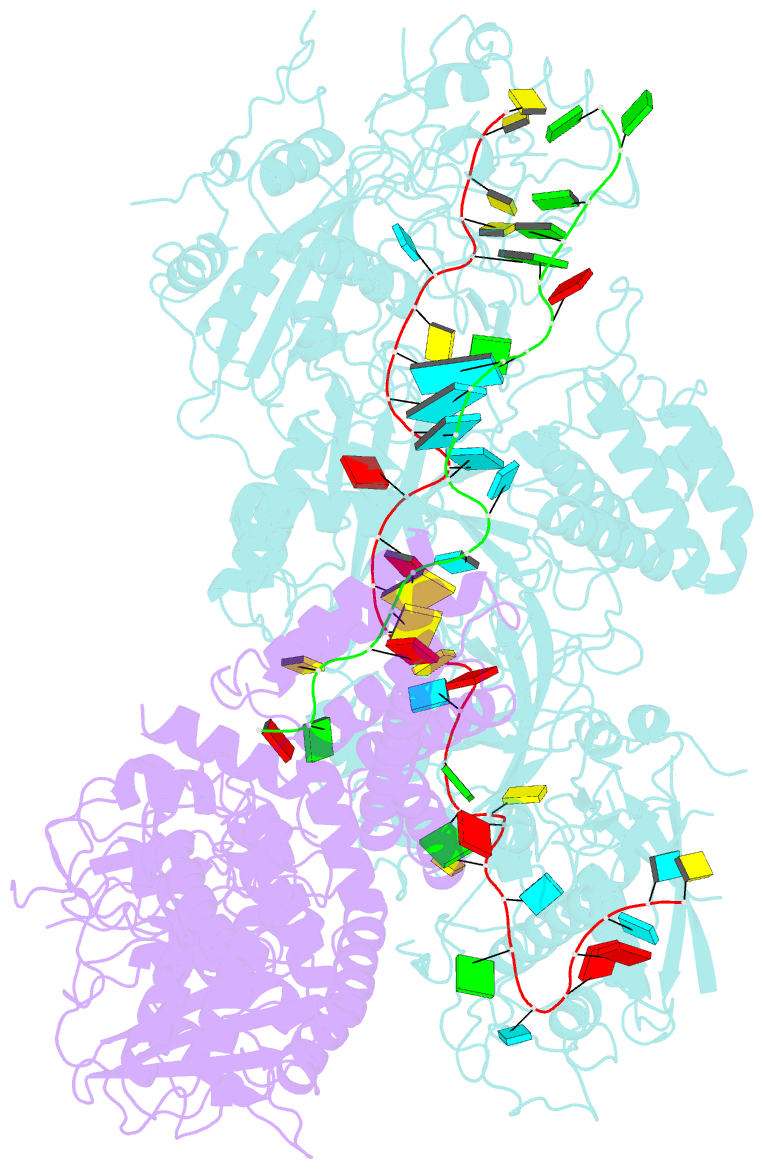
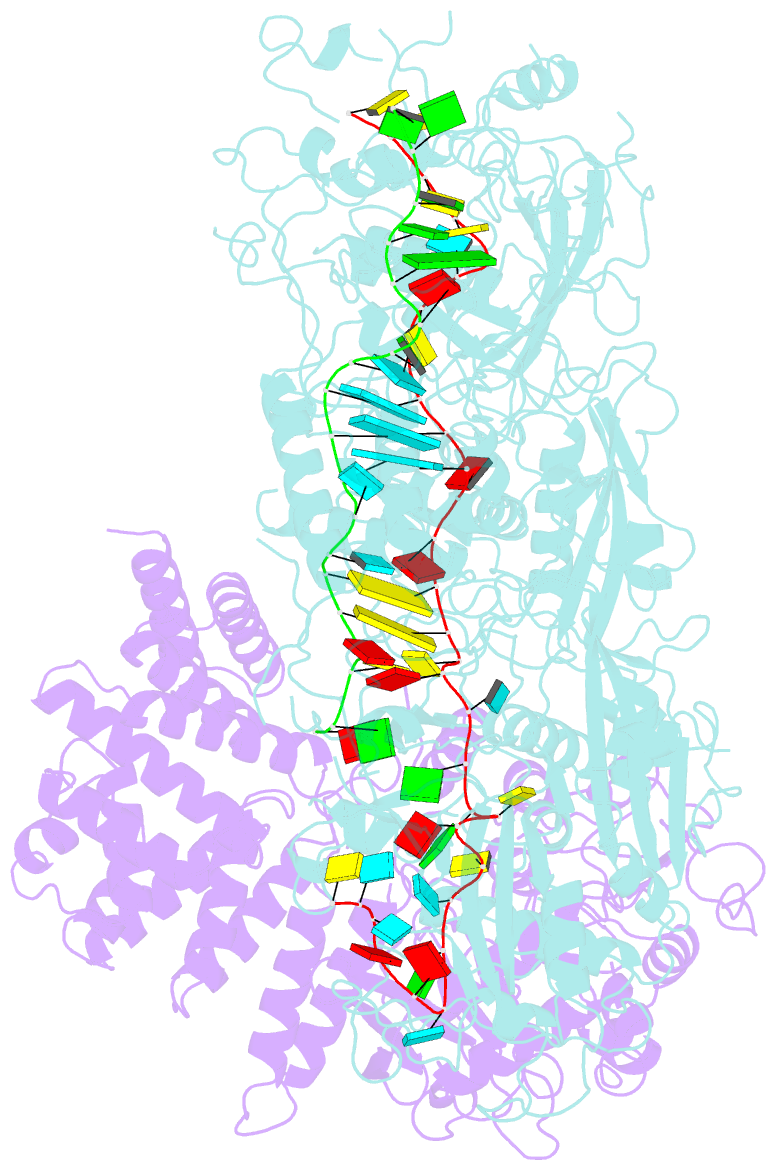
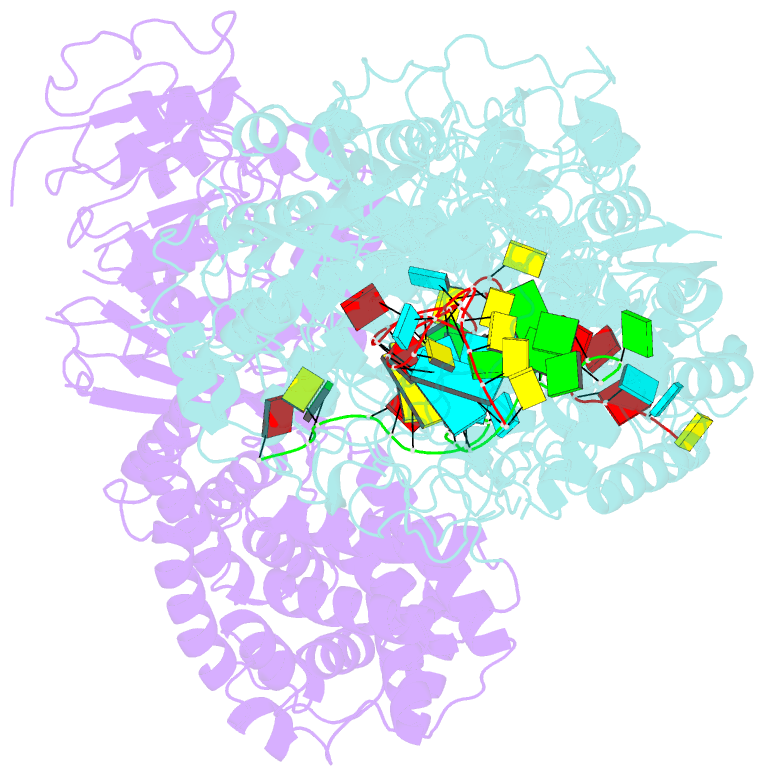
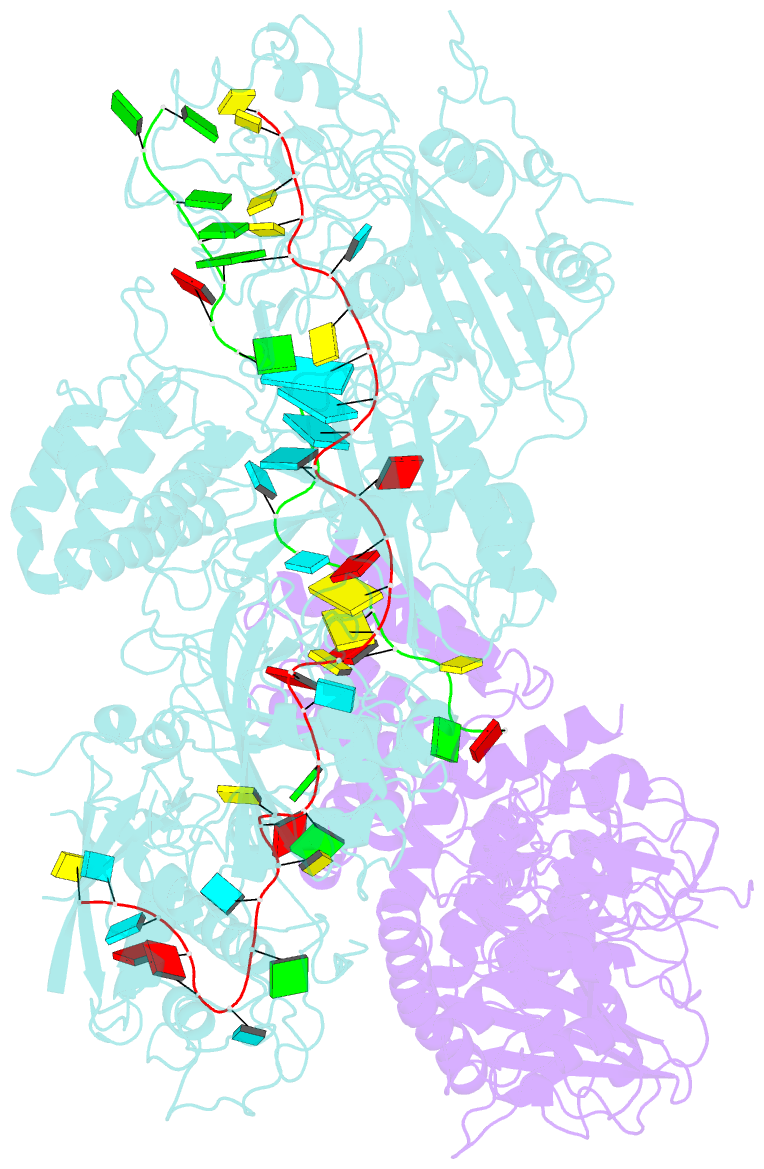
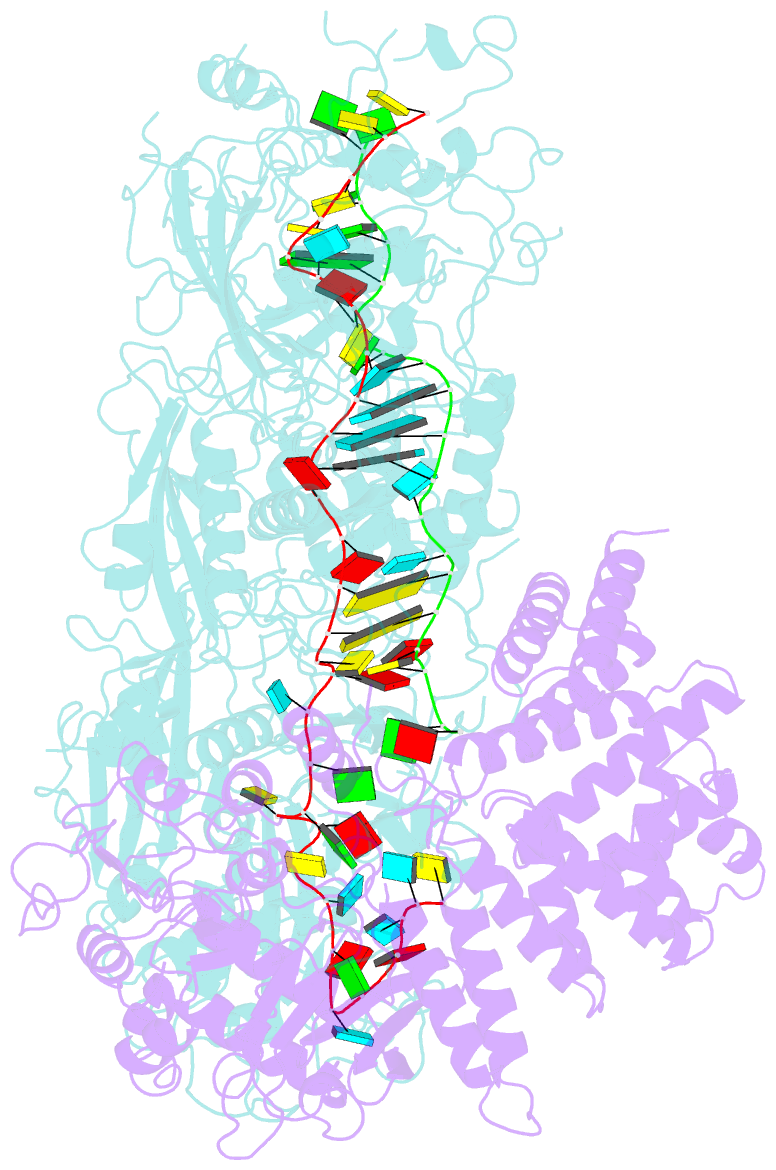
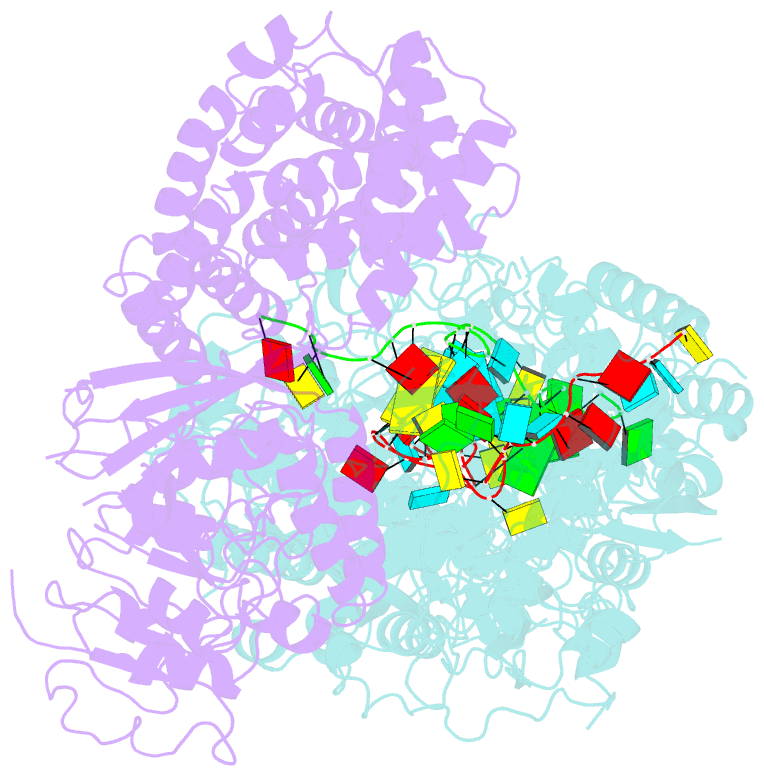
- cryo-EM structure of a bacterial protein complex
- Reference
- Yu G, Wang X, Zhang Y, An Q, Wen Y, Li X, Yin H, Deng Z, Zhang H (2022): "Structure and function of a bacterial type III-E CRISPR-Cas7-11 complex." Nat Microbiol, 7, 2078-2088. doi: 10.1038/s41564-022-01256-z.
- Abstract
- The type III-E CRISPR-Cas system uses a single multidomain effector called Cas7-11 (also named gRAMP) to cleave RNA and associate with a caspase-like protease Csx29, showing promising potential for RNA-targeting applications. The structural and molecular mechanisms of the type III-E CRISPR-Cas system remain poorly understood. Here we report four cryo-electron microscopy structures of Cas7-11 at different functional states. Cas7-11 has four Cas7-like domains, which assemble into a helical filament to accommodate CRISPR RNA (crRNA), and a Cas11-like domain facilitating crRNA-target RNA duplex formation. The Cas7.1 domain is critical for crRNA maturation, whereas Cas7.2 and Cas7.3 are responsible for target RNA cleavage. Target RNA binding induces the structural arrangements of Csx29, potentially exposing the catalytic site of Csx29. These results delineate the molecular mechanisms underlying pre-crRNA processing, target RNA recognition and cleavage for Cas7-11, and provide a structural framework to understand the role of Csx29 in type III-E CRISPR system.