Summary information and primary citation
- PDB-id
- 7xpg; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- virus like particle
- Method
- cryo-EM (3.51 Å)
- Summary
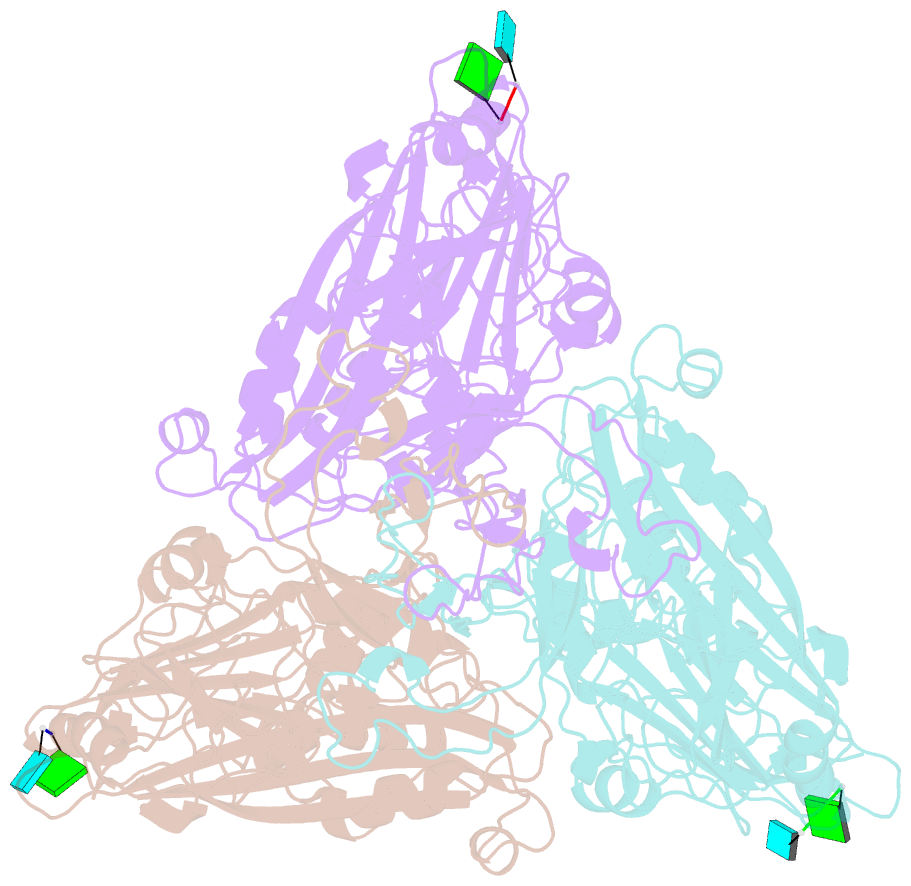
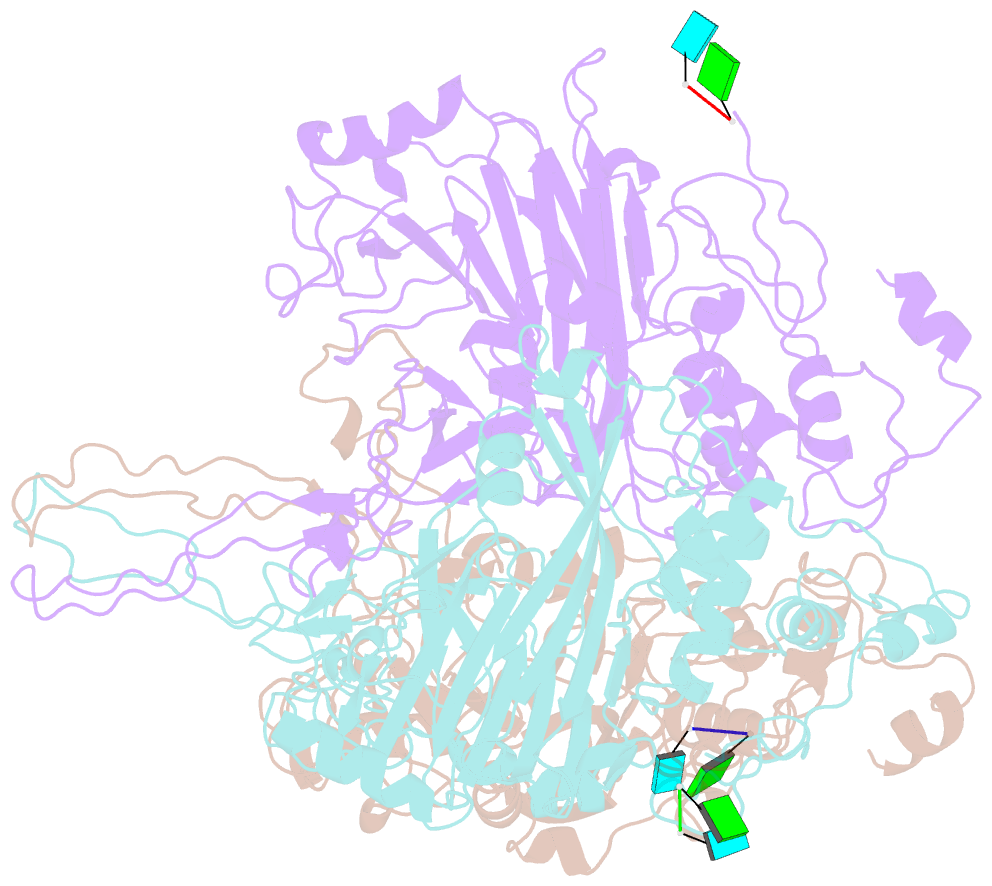
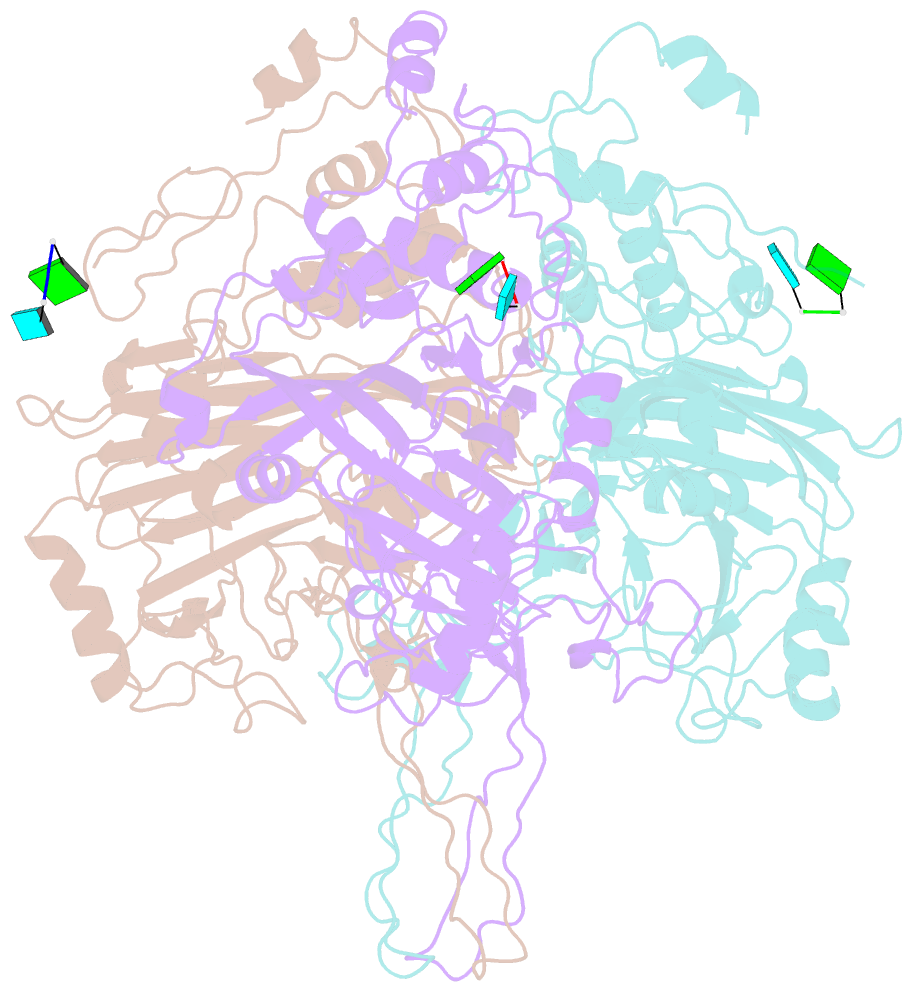
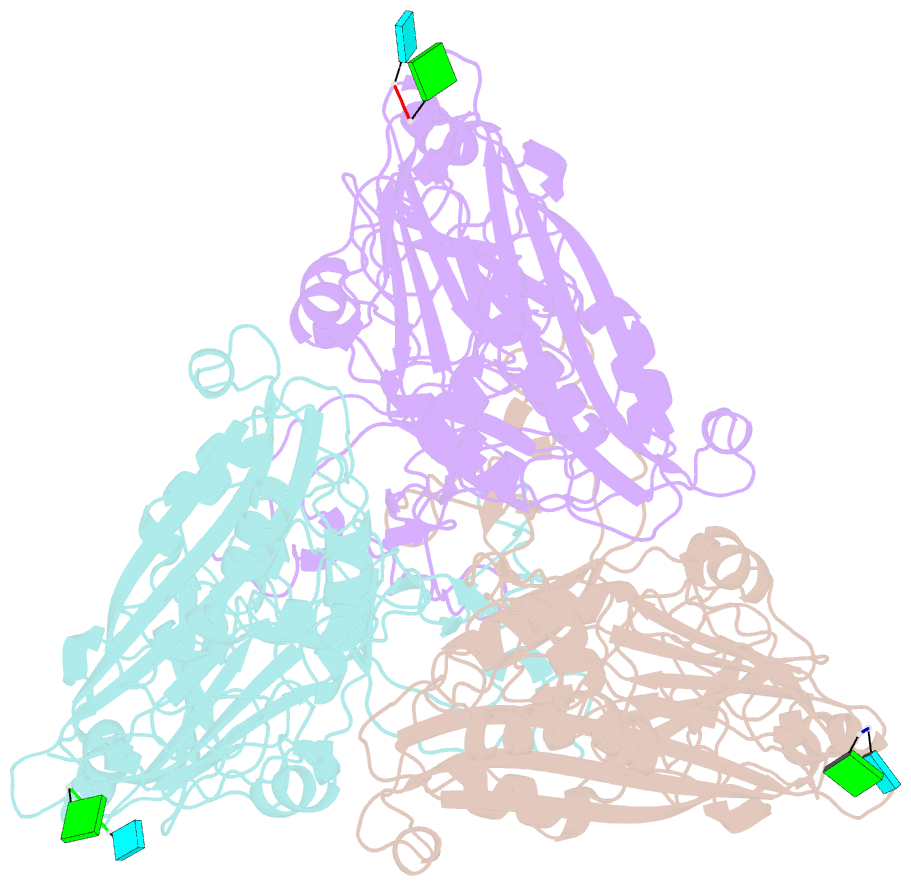
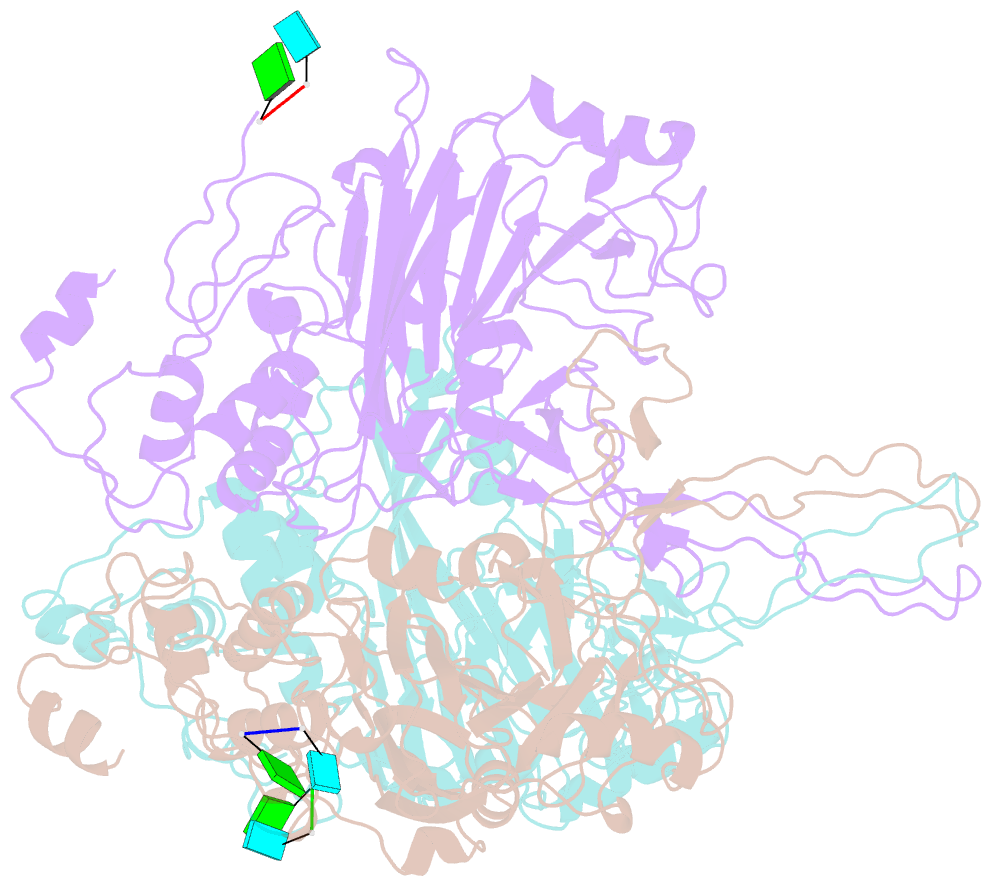
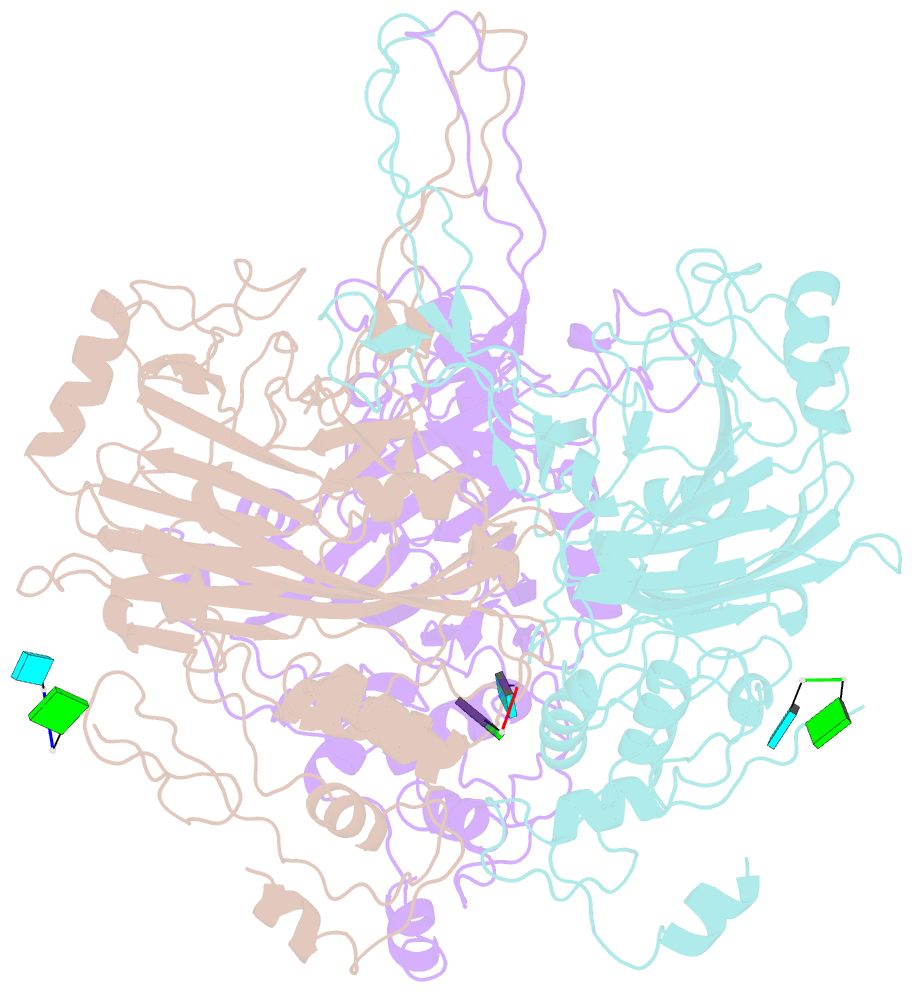
- cryo-EM structure of the t=3 lake sinai virus 1 (delta-n48) virus-like capsid at ph 6.5
- Reference
- Chen NC, Wang CH, Yoshimura M, Yeh YQ, Guan HH, Chuankhayan P, Lin CC, Lin PJ, Huang YC, Wakatsuki S, Ho MC, Chen CJ (2023): "Structures of honeybee-infecting Lake Sinai virus reveal domain functions and capsid assembly with dynamic motions." Nat Commun, 14, 545. doi: 10.1038/s41467-023-36235-3.
- Abstract
- Understanding the structural diversity of honeybee-infecting viruses is critical to maintain pollinator health and manage the spread of diseases in ecology and agriculture. We determine cryo-EM structures of T = 4 and T = 3 capsids of virus-like particles (VLPs) of Lake Sinai virus (LSV) 2 and delta-N48 LSV1, belonging to tetraviruses, at resolutions of 2.3-2.6 Å in various pH environments. Structural analysis shows that the LSV2 capsid protein (CP) structural features, particularly the protruding domain and C-arm, differ from those of other tetraviruses. The anchor loop on the central β-barrel domain interacts with the neighboring subunit to stabilize homo-trimeric capsomeres during assembly. Delta-N48 LSV1 CP interacts with ssRNA via the rigid helix α1', α1'-α1 loop, β-barrel domain, and C-arm. Cryo-EM reconstructions, combined with X-ray crystallographic and small-angle scattering analyses, indicate that pH affects capsid conformations by regulating reversible dynamic particle motions and sizes of LSV2 VLPs. C-arms exist in all LSV2 and delta-N48 LSV1 VLPs across varied pH conditions, indicating that autoproteolysis cleavage is not required for LSV maturation. The observed linear domino-scaffold structures of various lengths, made up of trapezoid-shape capsomeres, provide a basis for icosahedral T = 4 and T = 3 architecture assemblies. These findings advance understanding of honeybee-infecting viruses that can cause Colony Collapse Disorder.