Summary information and primary citation
- PDB-id
- 7y7q; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- RNA binding protein
- Method
- X-ray (2.05 Å)
- Summary
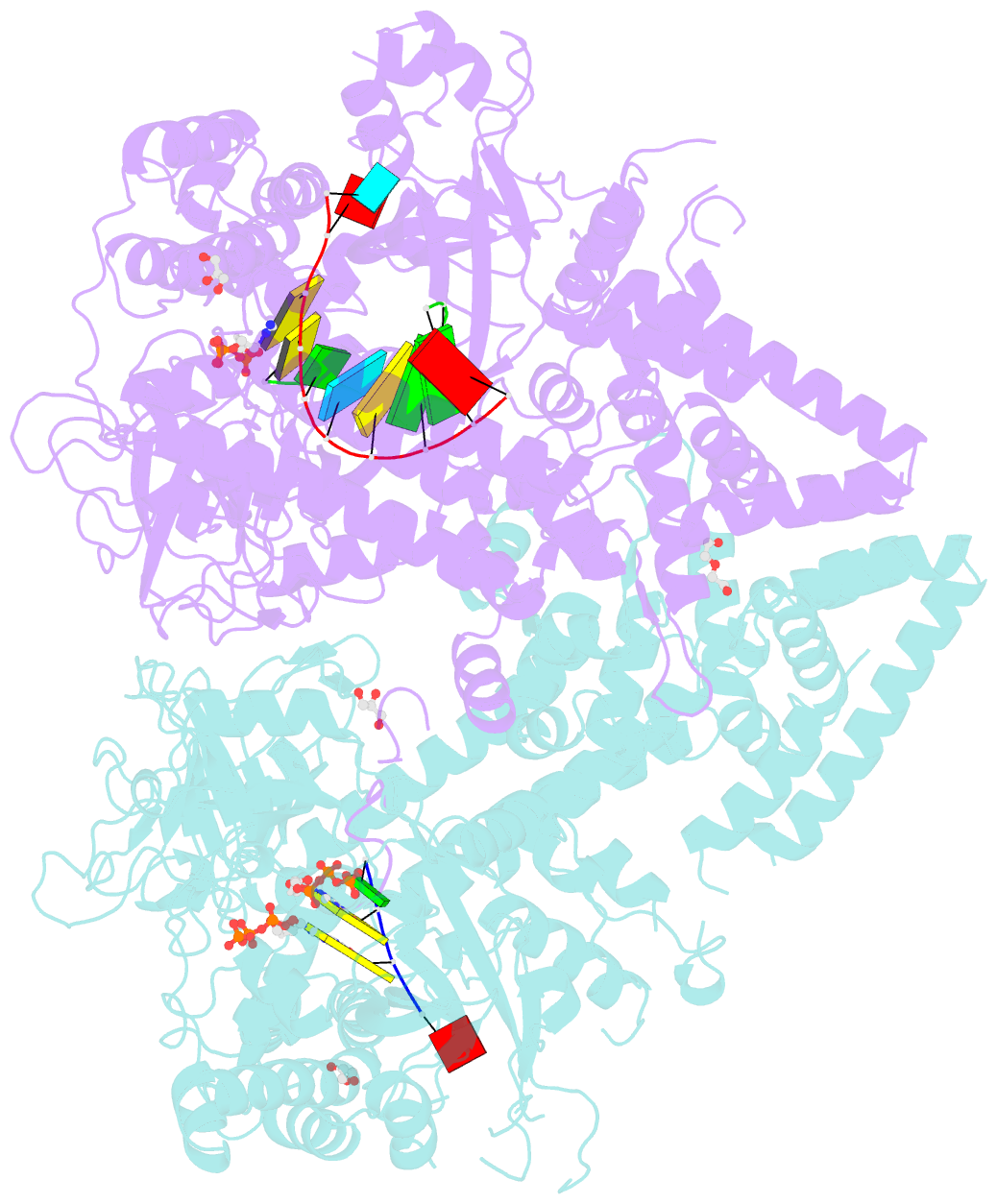
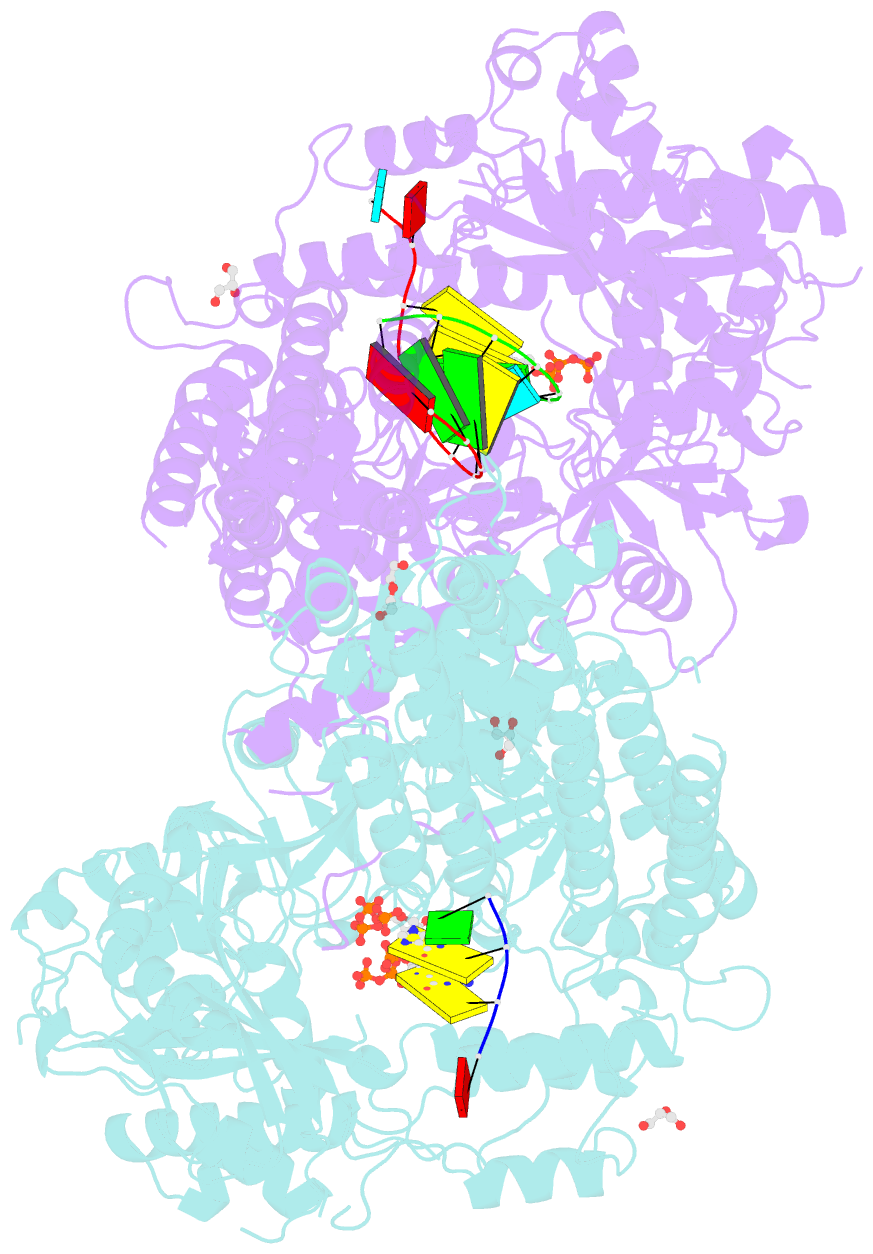
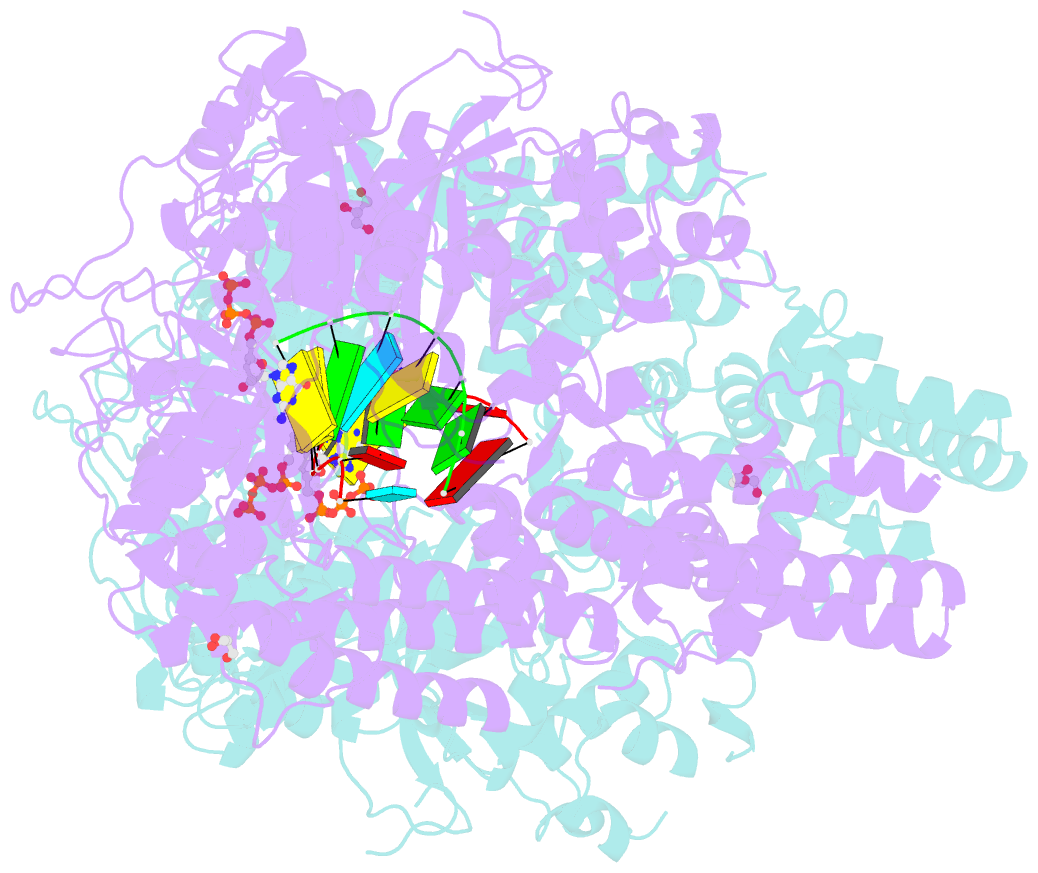
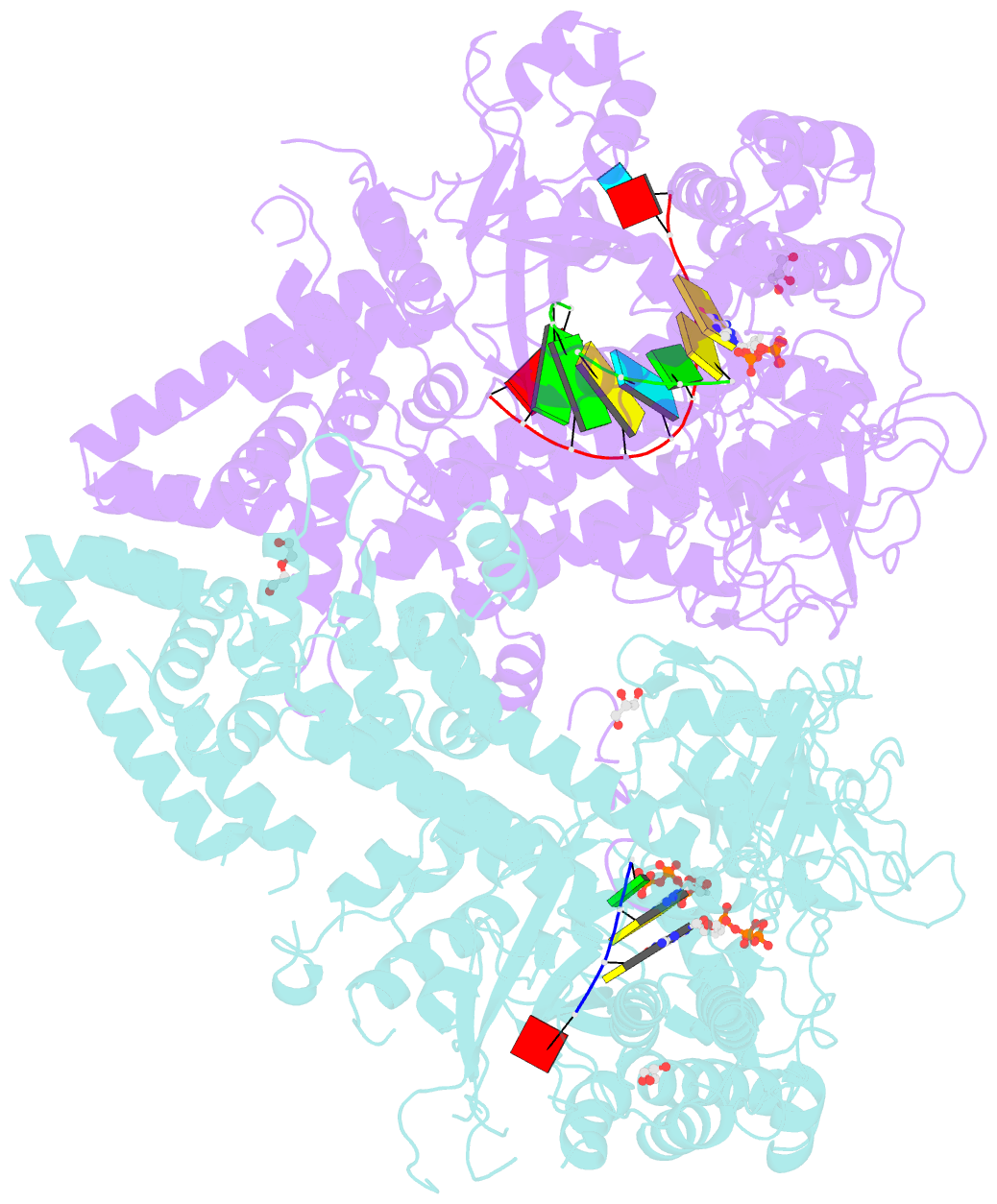
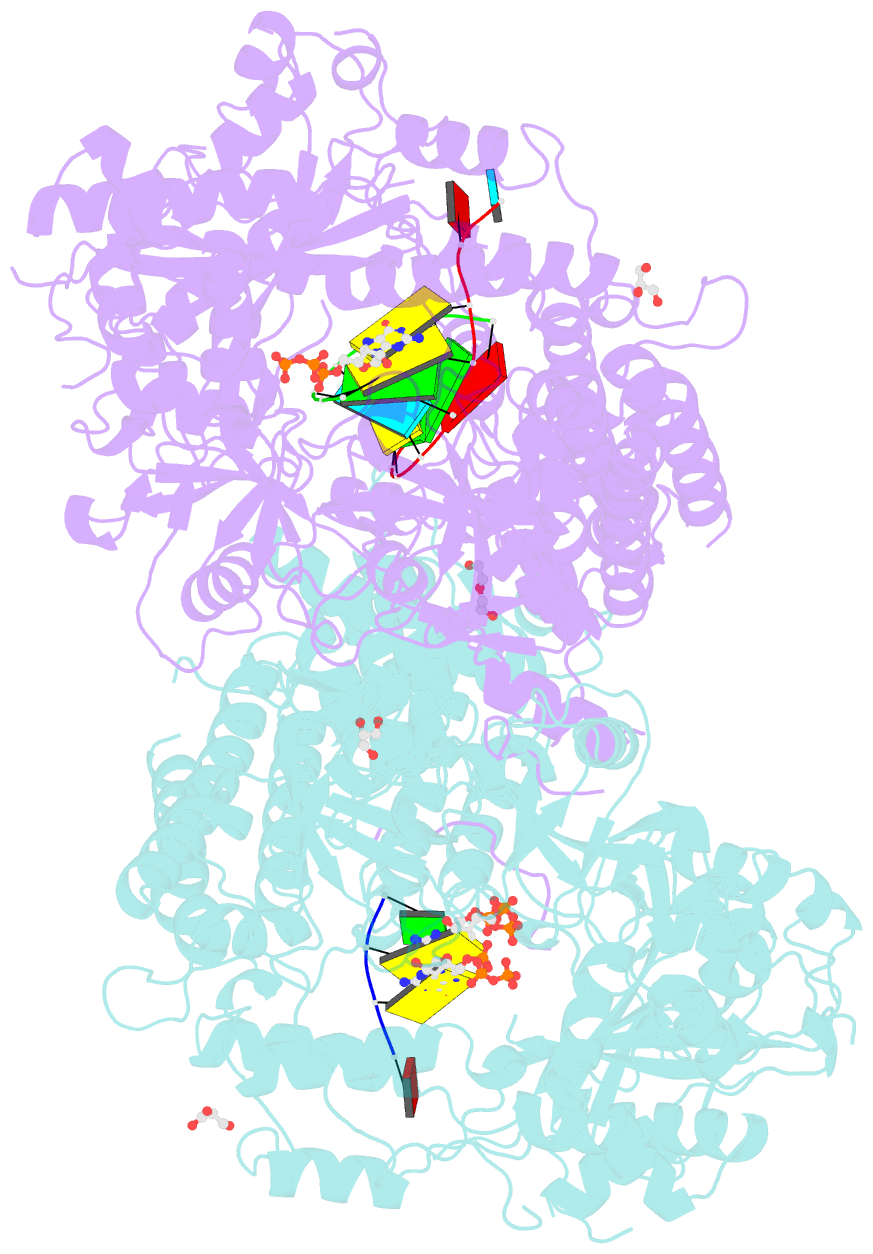
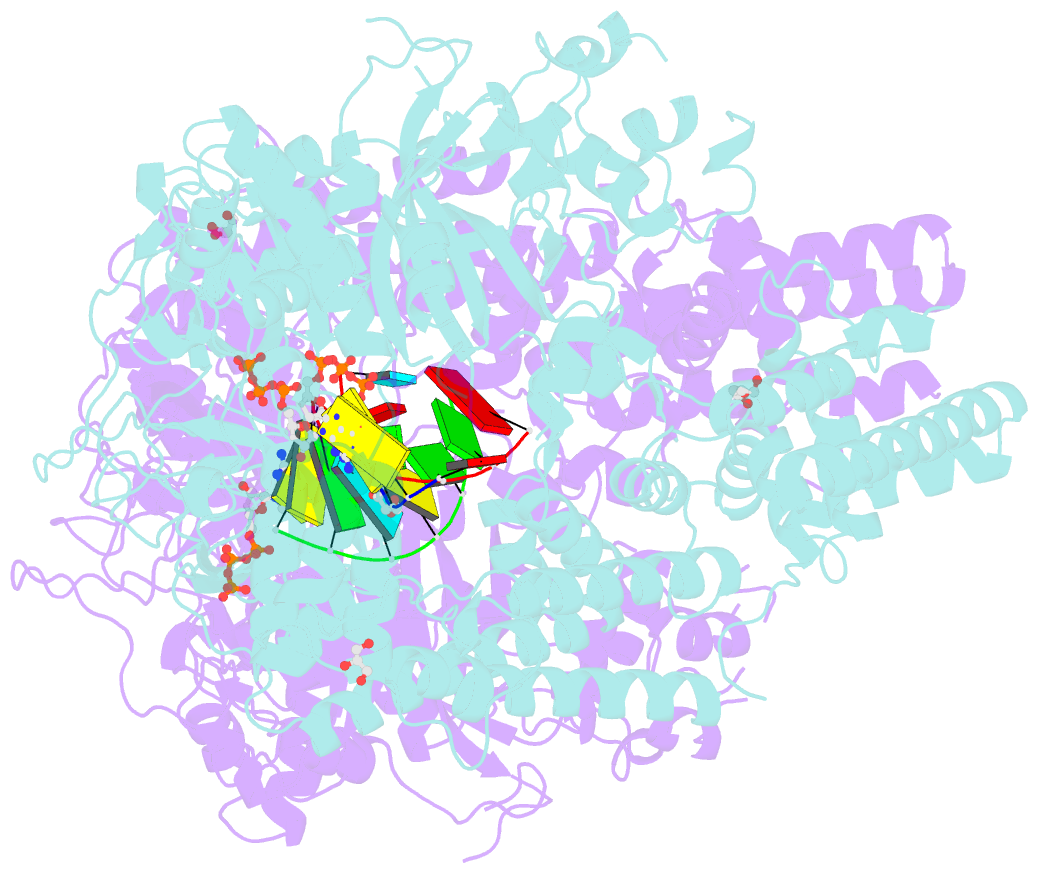
- Qde-1 in complex with RNA template, RNA primer and 3'-dgtp
- Reference
- Cui R, Li H, Zhao J, Li X, Gan J, Ma J (2022): "Structural insights into the dual activities of the two-barrel RNA polymerase QDE-1." Nucleic Acids Res., 50, 10169-10186. doi: 10.1093/nar/gkac727.
- Abstract
- Neurospora crassa protein QDE-1, a member of the two-barrel polymerase superfamily, possesses both DNA- and RNA-dependent RNA polymerase (DdRP and RdRP) activities. The dual activities are essential for the production of double-stranded RNAs (dsRNAs), the precursors of small interfering RNAs (siRNAs) in N. crassa. Here, we report five complex structures of N-terminal truncated QDE-1 (QDE-1ΔN), representing four different reaction states: DNA/RNA-templated elongation, the de novo initiation of RNA synthesis, the first step of nucleotide condensation during de novo initiation and initial NTP loading. The template strand is aligned by a bridge-helix and double-psi beta-barrels 2 (DPBB2), the RNA product is held by DPBB1 and the slab domain. The DNA template unpairs with the RNA product at position -7, but the RNA template remains paired. The NTP analog coordinates with cations and is precisely positioned at the addition site by a rigid trigger loop and a proline-containing loop in the active center. The unique C-terminal tail from the QDE-1 dimer partner inserts into the substrate-binding cleft and plays regulatory roles in RNA synthesis. Collectively, this work elucidates the conserved mechanisms for DNA/RNA-dependent dual activities by QDE-1 and other two-barrel polymerase superfamily members.