Summary information and primary citation
- PDB-id
- 7yza; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- DNA binding protein
- Method
- X-ray (1.18 Å)
- Summary
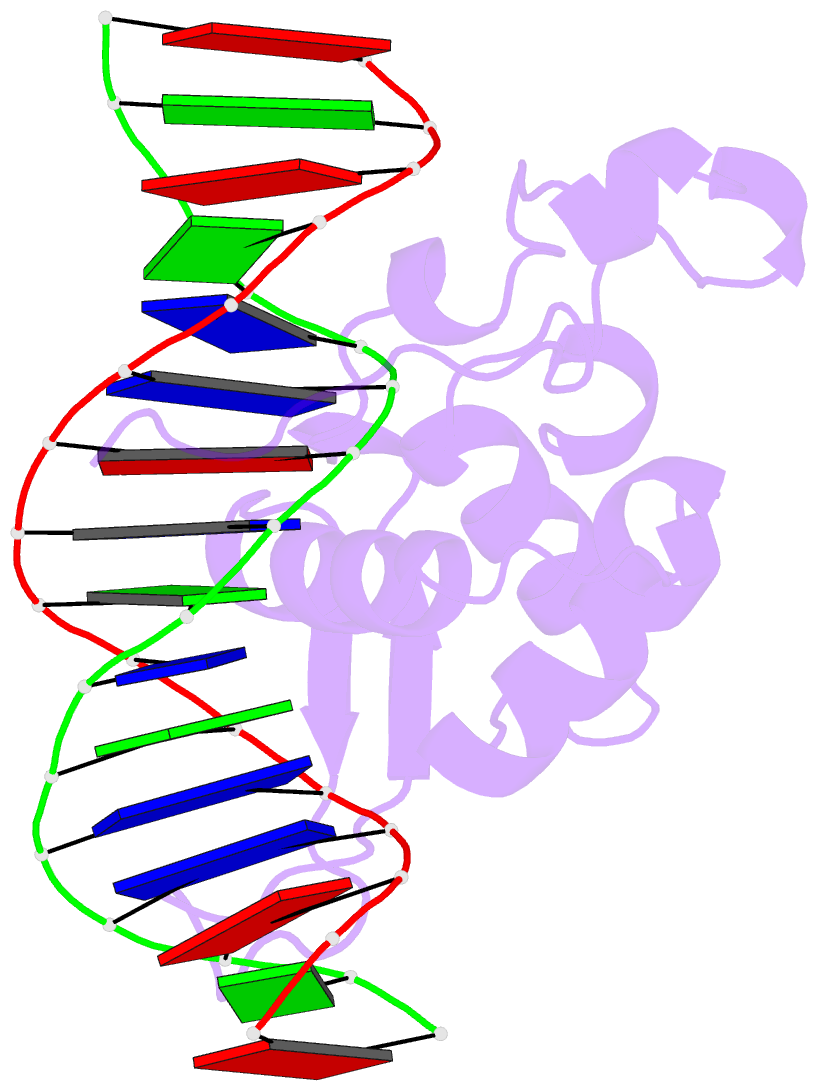
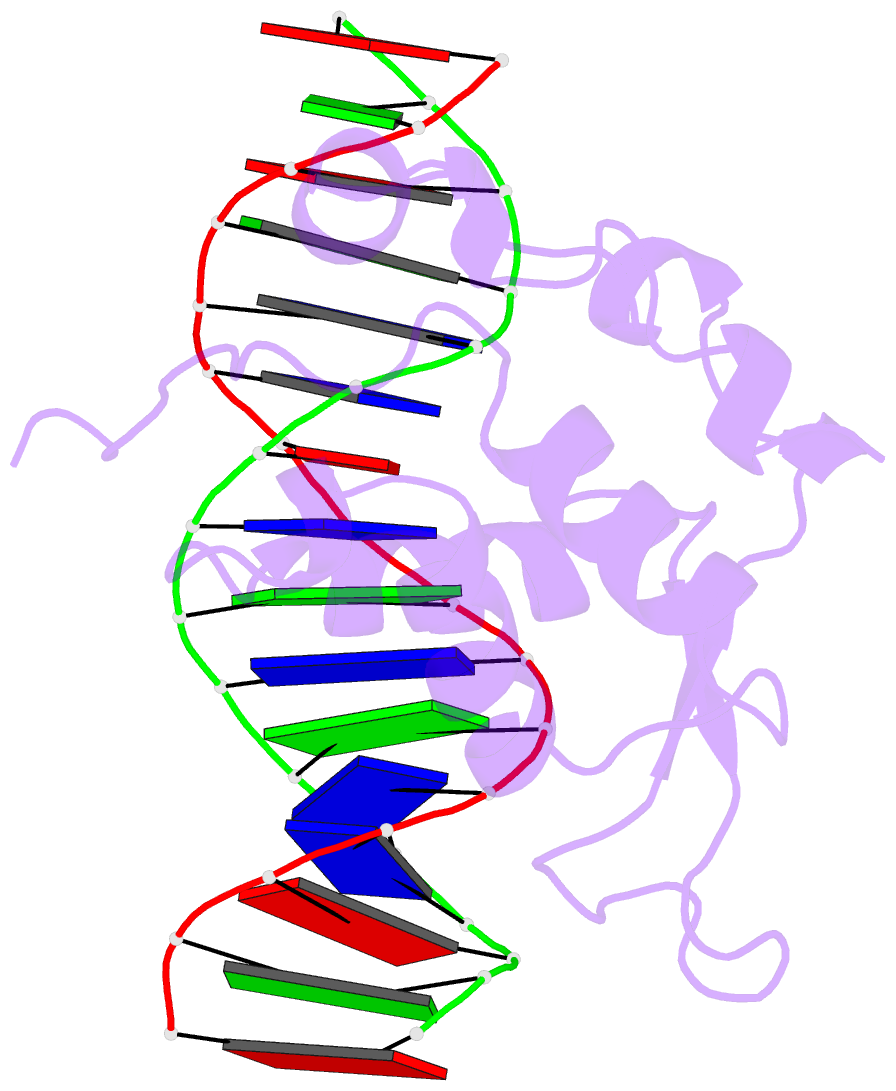
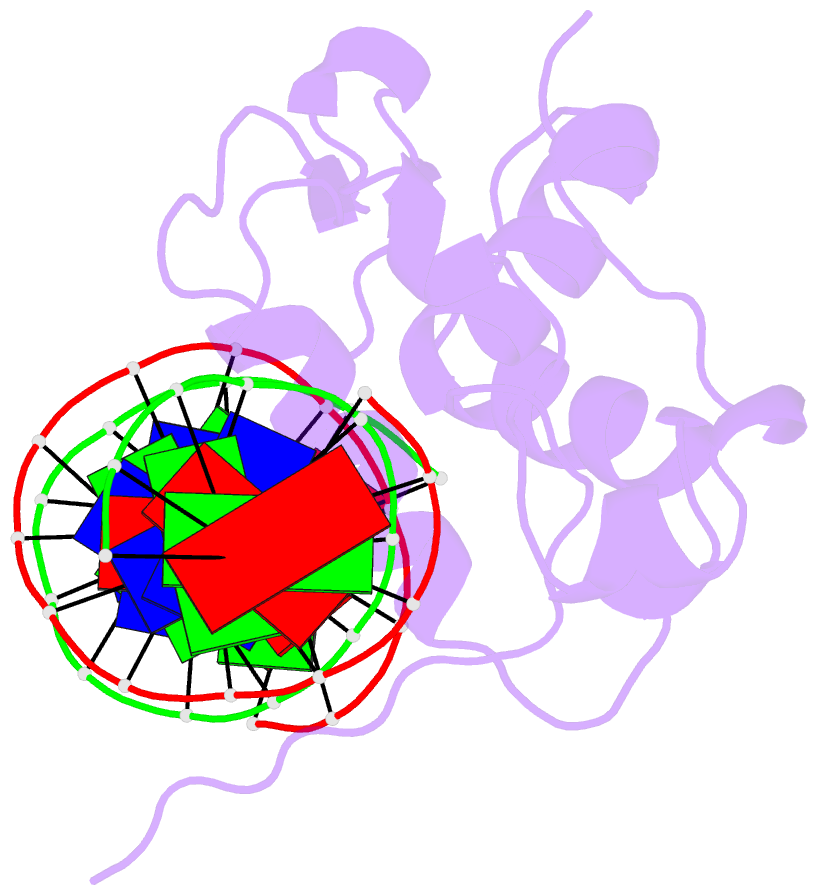
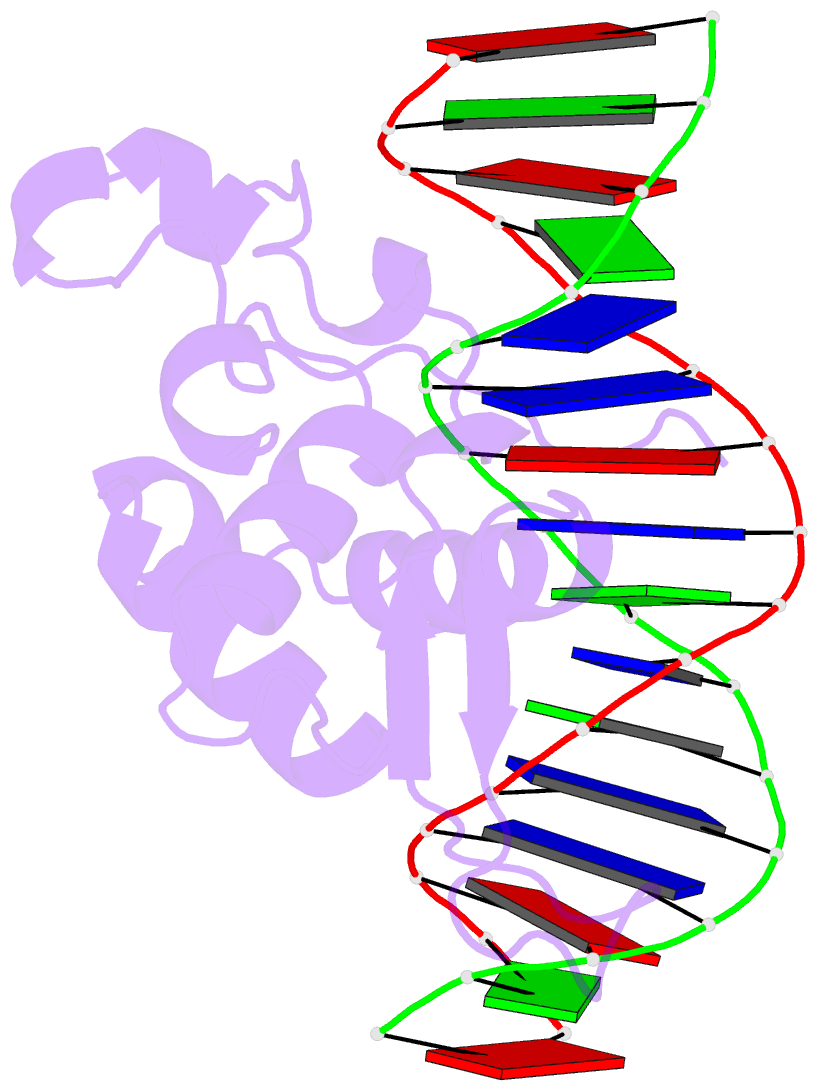
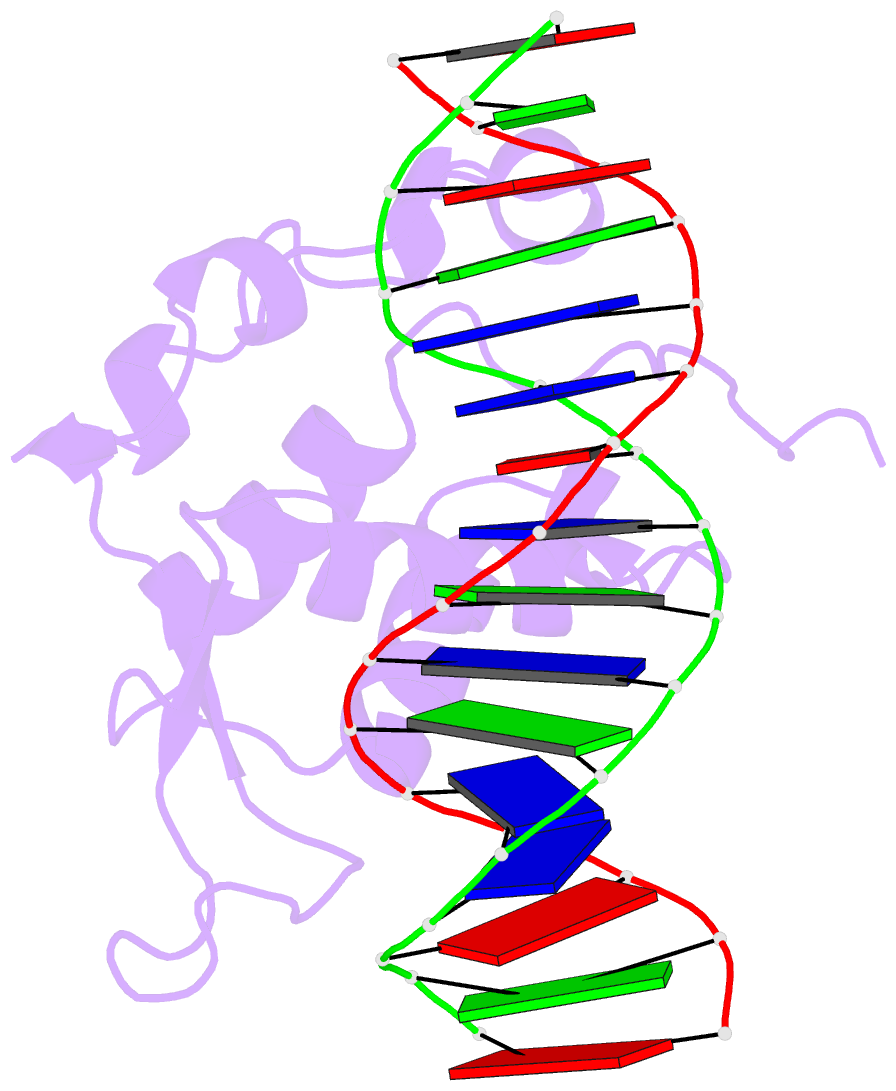
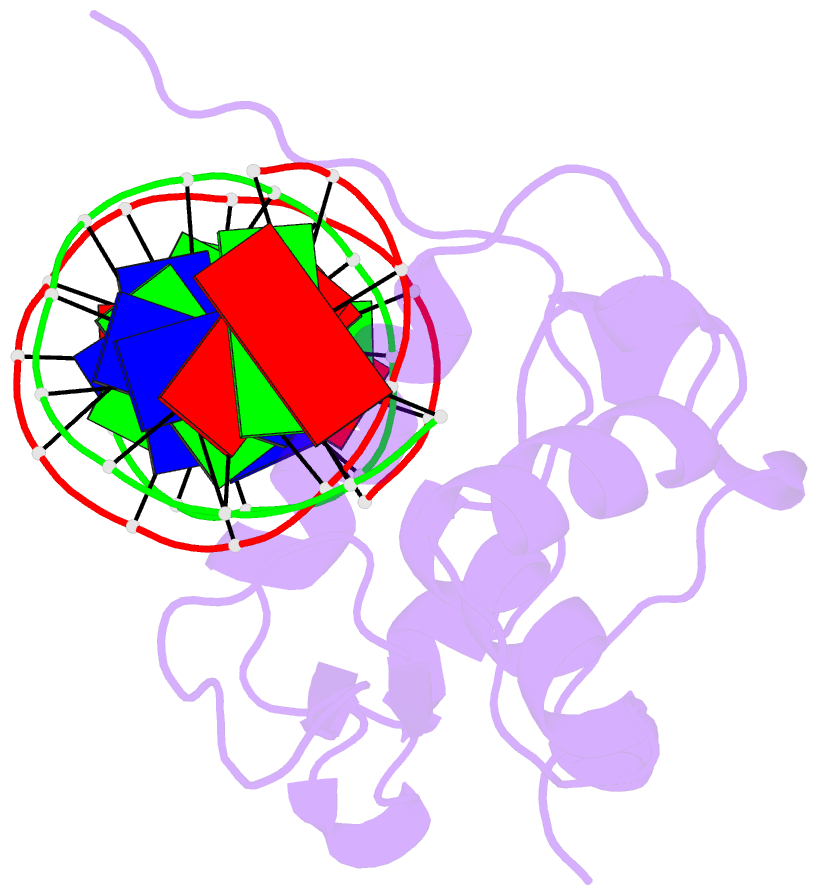
- Crystal structure of the zebrafish foxh1 bound to the tgtgtatt site
- Reference
- Pluta R, Aragon E, Prescott NA, Ruiz L, Mees RA, Baginski B, Flood JR, Martin-Malpartida P, Massague J, David Y, Macias MJ (2022): "Molecular basis for DNA recognition by the maternal pioneer transcription factor FoxH1." Nat Commun, 13, 7279. doi: 10.1038/s41467-022-34925-y.
- Abstract
- Forkhead box H1 (FoxH1) is an essential maternal pioneer factor during embryonic development that binds to specific GG/GT-containing DNA target sequences. Here we have determined high-resolution structures of three FoxH1 proteins (from human, frog and fish species) and four DNAs to clarify the way in which FoxH1 binds to these sites. We found that the protein-DNA interactions extend to both the minor and major DNA grooves and are thus almost twice as extensive as those of other FOX family members. Moreover, we identified two specific amino acid changes in FoxH1 that allowed the recognition of GG/GT motifs. Consistent with the pioneer factor activity of FoxH1, we found that its affinity for nucleosomal DNA is even higher than for linear DNA fragments. The structures reported herein illustrate how FoxH1 binding to distinct DNA sites provides specificity and avoids cross-regulation by other FOX proteins that also operate during the maternal-zygotic transition and select canonical forkhead sites.