Summary information and primary citation
- PDB-id
- 7zpk; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- RNA binding protein
- Method
- cryo-EM (3.81 Å)
- Summary
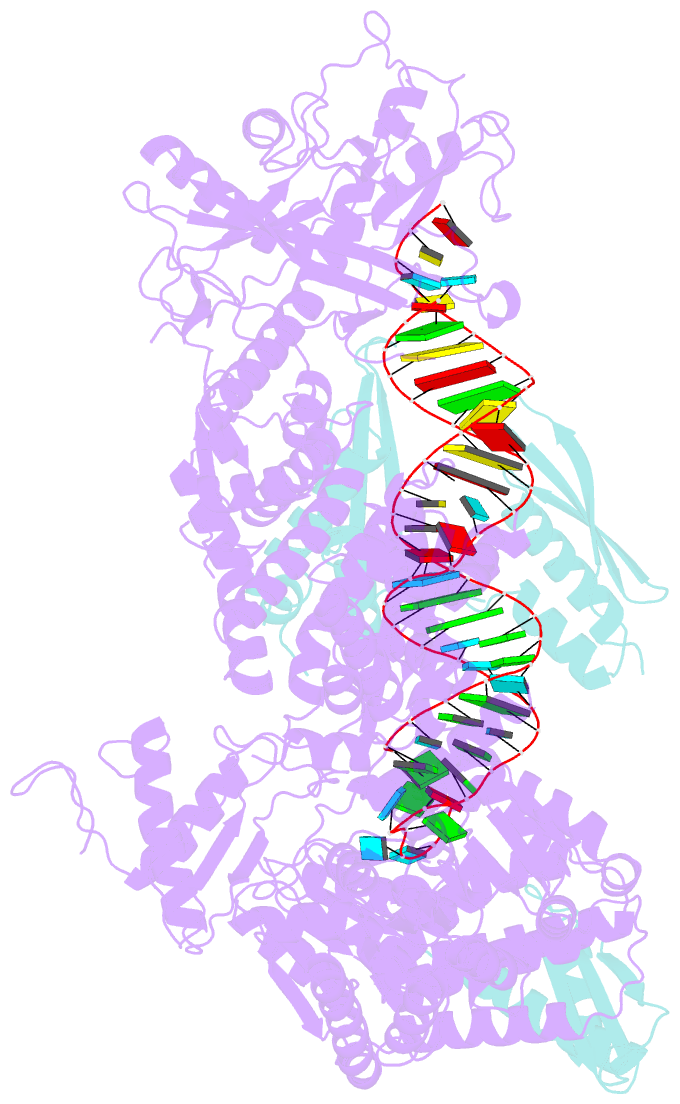
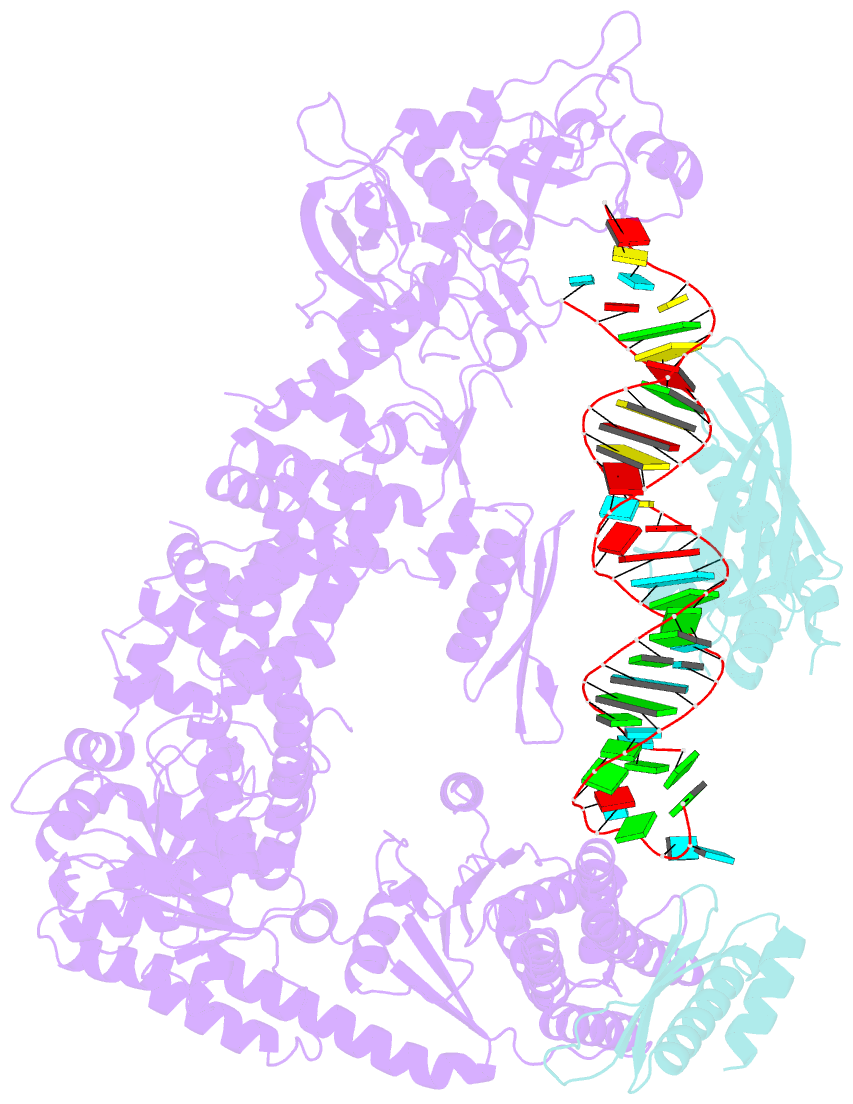
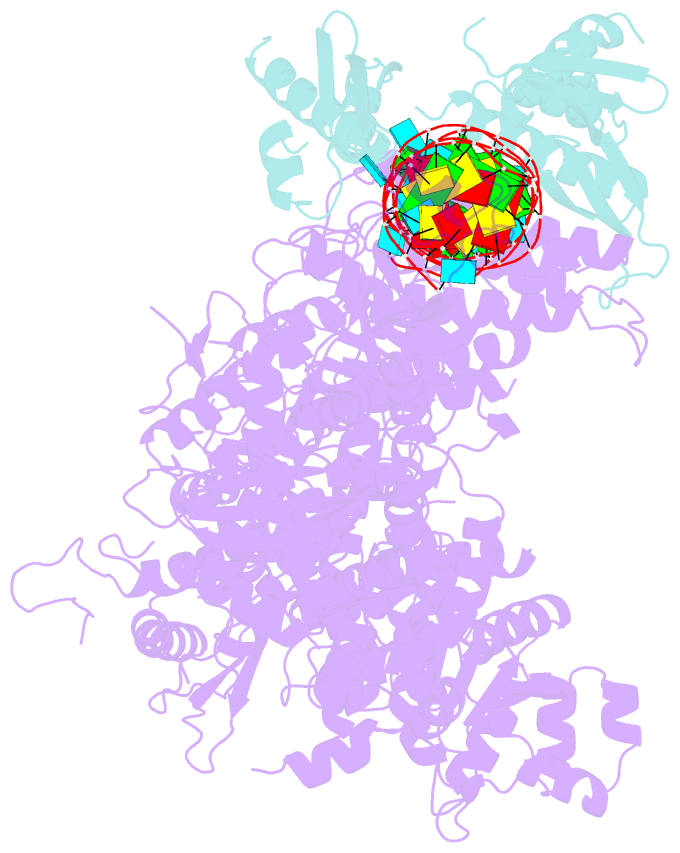
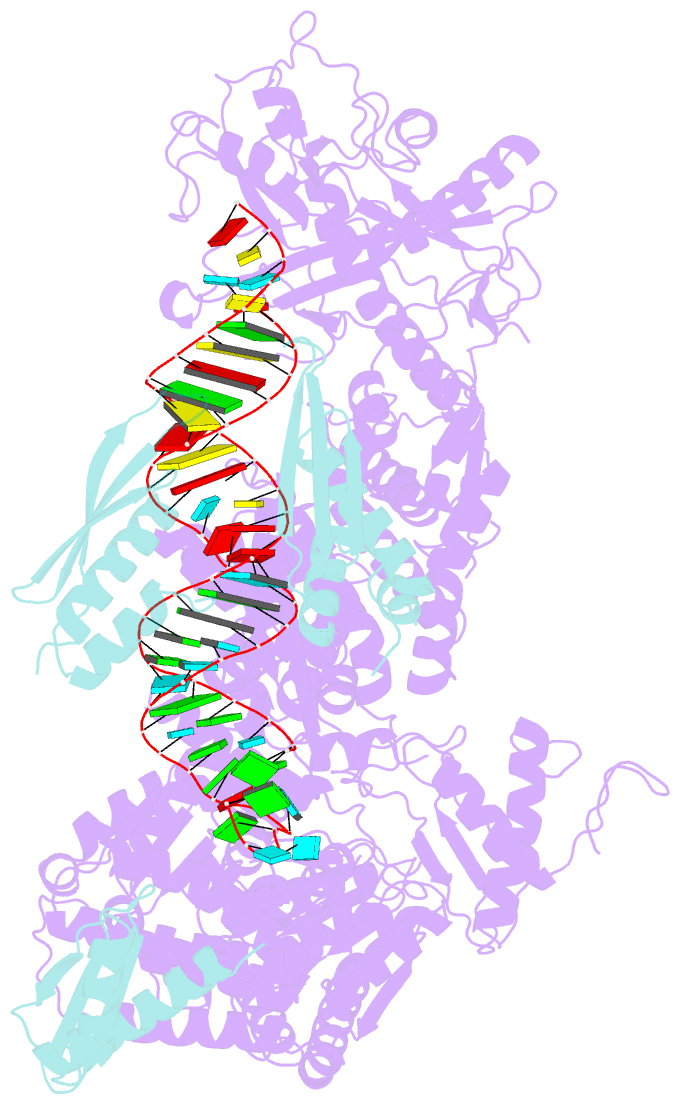
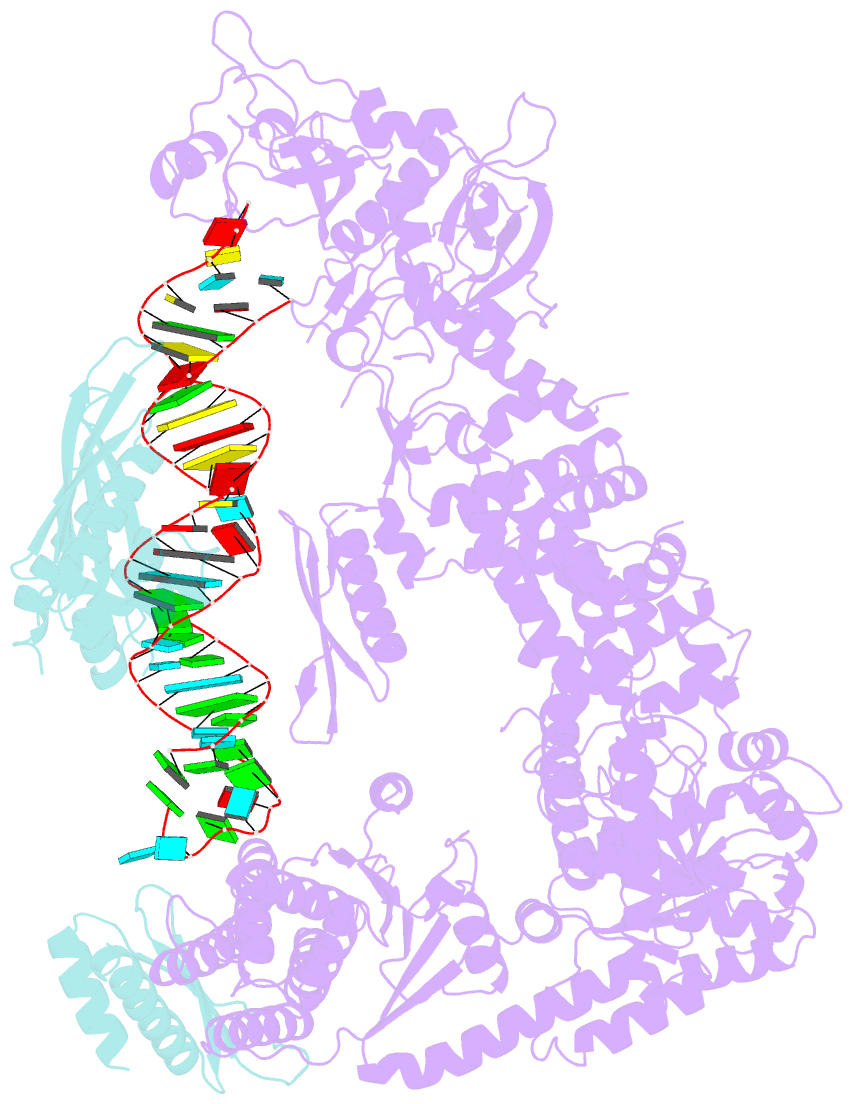
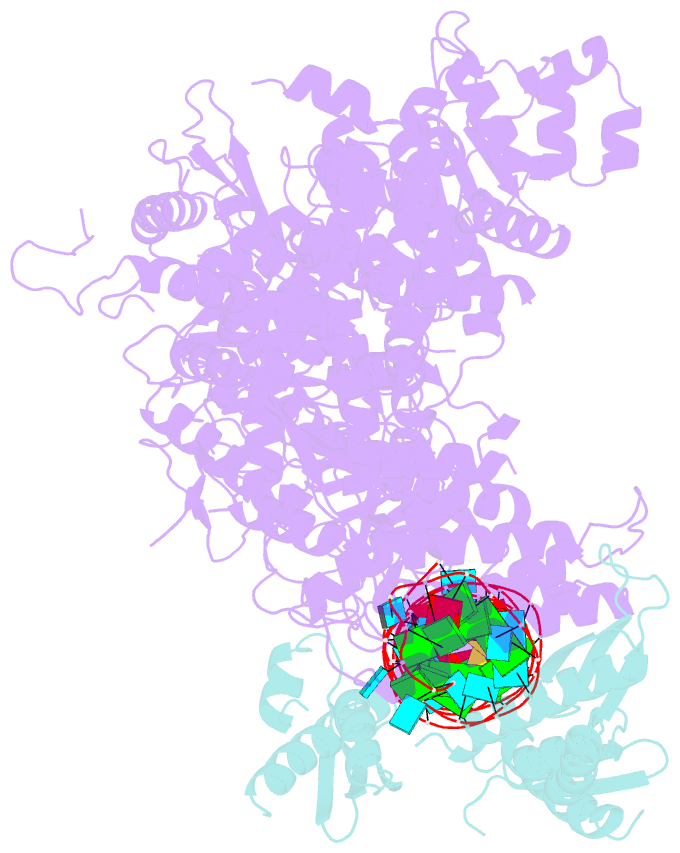
- Mammalian dicer in the "pre-dicing state" with pre-mir-15a substrate and tarbp2 subunit
- Reference
- Zapletal D, Taborska E, Pasulka J, Malik R, Kubicek K, Zanova M, Much C, Sebesta M, Buccheri V, Horvat F, Jenickova I, Prochazkova M, Prochazka J, Pinkas M, Novacek J, Joseph DF, Sedlacek R, Bernecky C, O'Carroll D, Stefl R, Svoboda P (2022): "Structural and functional basis of mammalian microRNA biogenesis by Dicer." Mol.Cell, 82, 4064-4079.e13. doi: 10.1016/j.molcel.2022.10.010.
- Abstract
- MicroRNA (miRNA) and RNA interference (RNAi) pathways rely on small RNAs produced by Dicer endonucleases. Mammalian Dicer primarily supports the essential gene-regulating miRNA pathway, but how it is specifically adapted to miRNA biogenesis is unknown. We show that the adaptation entails a unique structural role of Dicer's DExD/H helicase domain. Although mice tolerate loss of its putative ATPase function, the complete absence of the domain is lethal because it assures high-fidelity miRNA biogenesis. Structures of murine Dicer•-miRNA precursor complexes revealed that the DExD/H domain has a helicase-unrelated structural function. It locks Dicer in a closed state, which facilitates miRNA precursor selection. Transition to a cleavage-competent open state is stimulated by Dicer-binding protein TARBP2. Absence of the DExD/H domain or its mutations unlocks the closed state, reduces substrate selectivity, and activates RNAi. Thus, the DExD/H domain structurally contributes to mammalian miRNA biogenesis and underlies mechanistical partitioning of miRNA and RNAi pathways.