Summary information and primary citation
- PDB-id
- 8ac8; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- DNA binding protein
- Method
- X-ray (1.6 Å)
- Summary
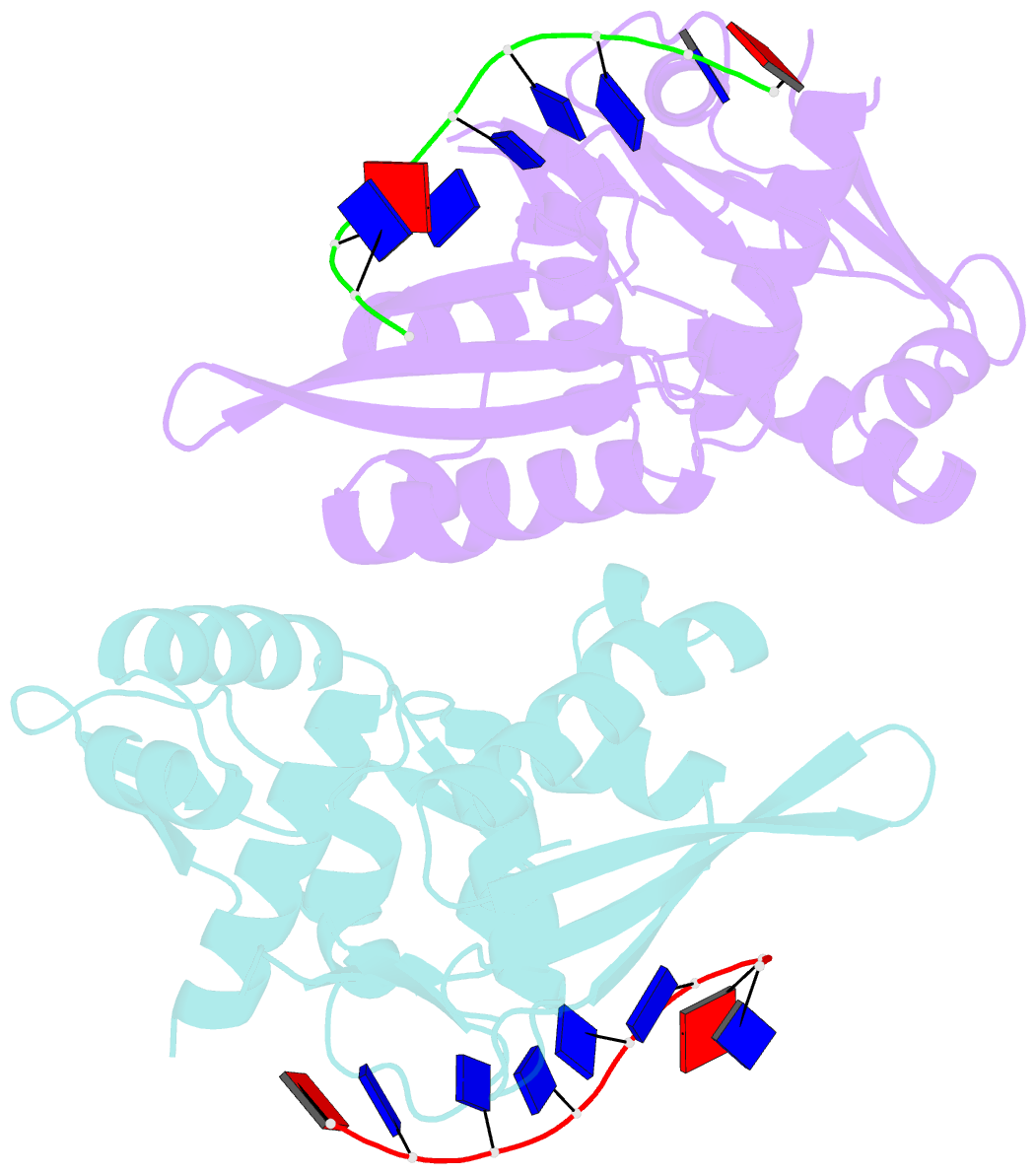
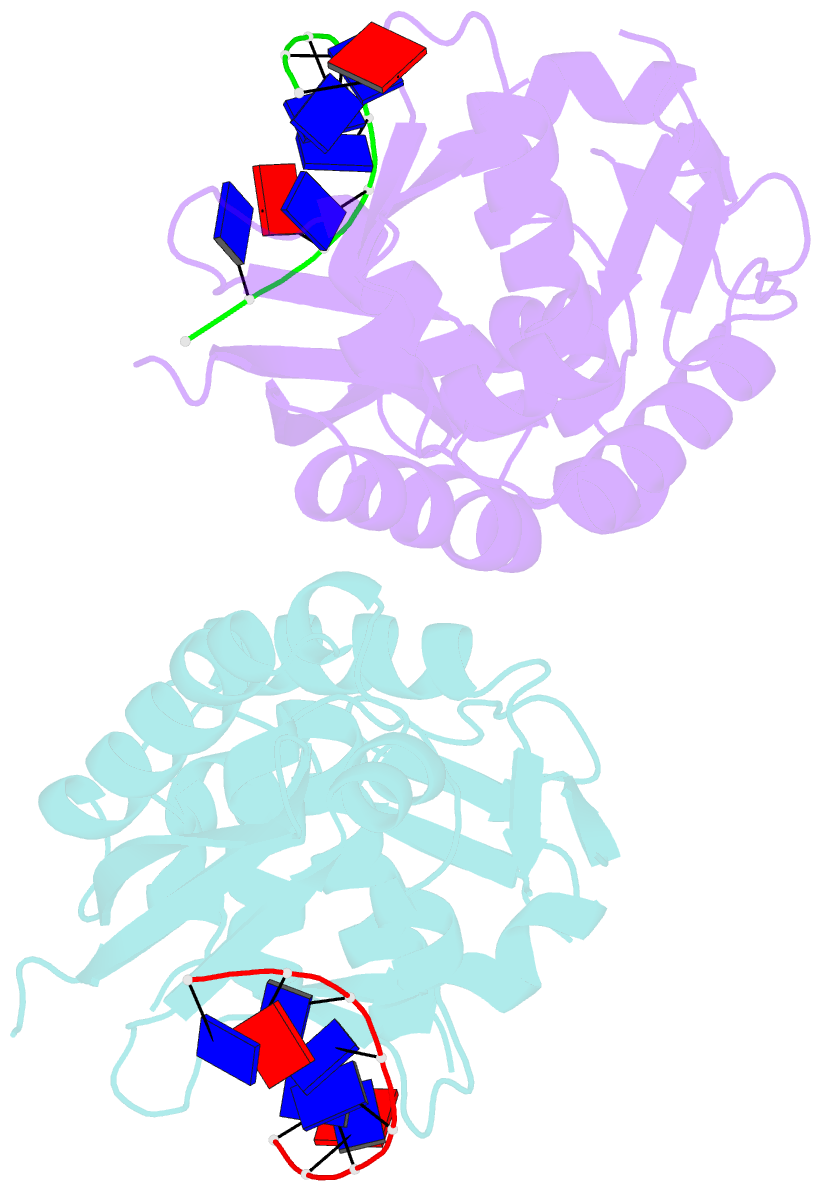
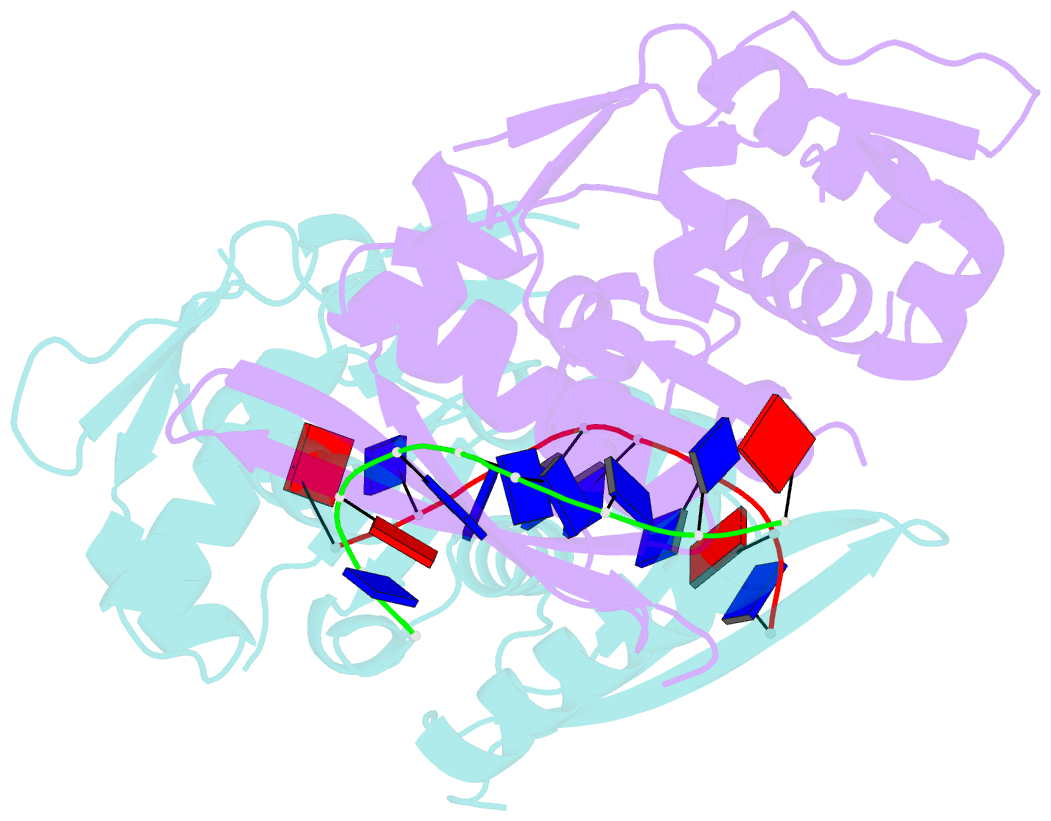
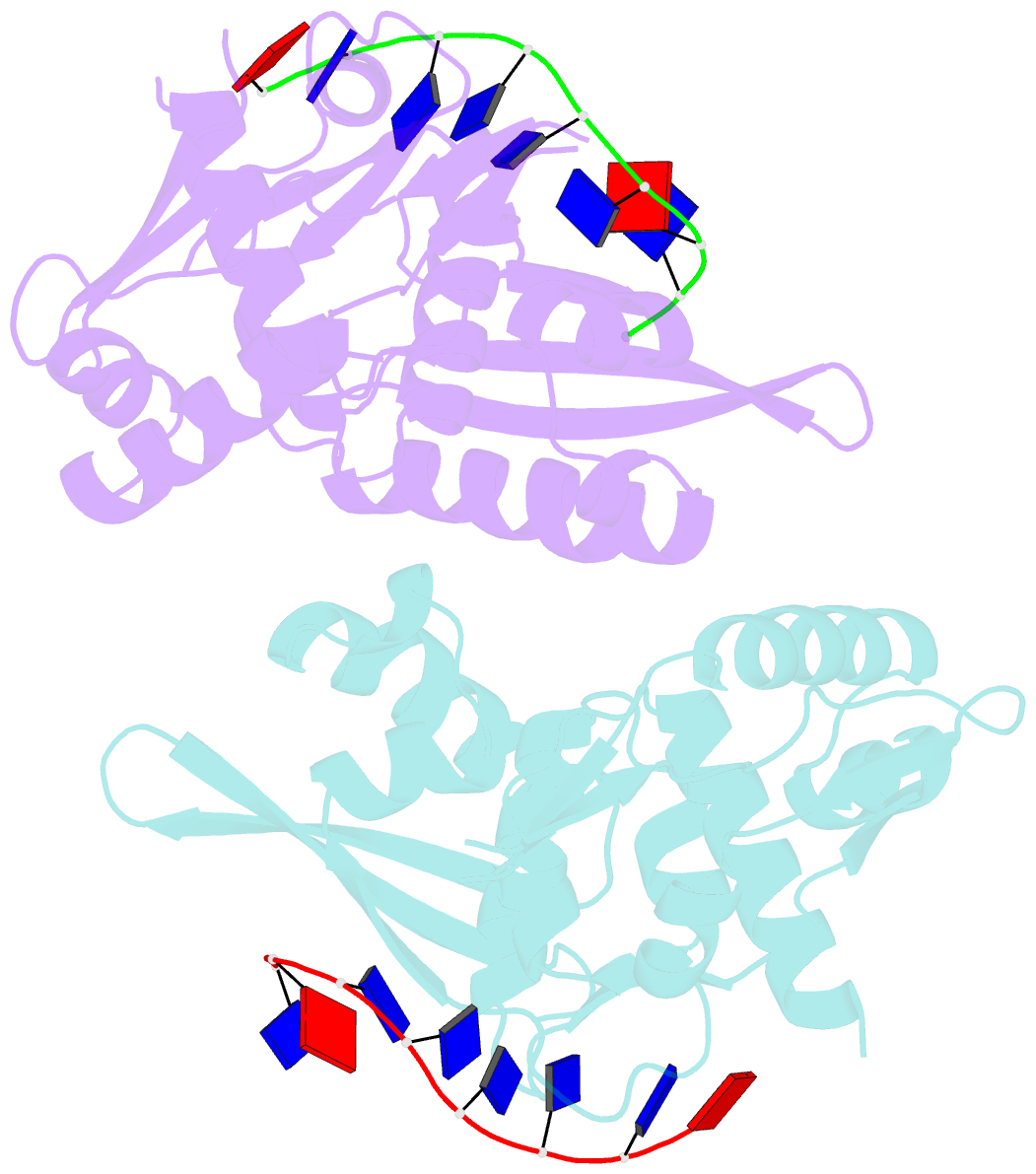
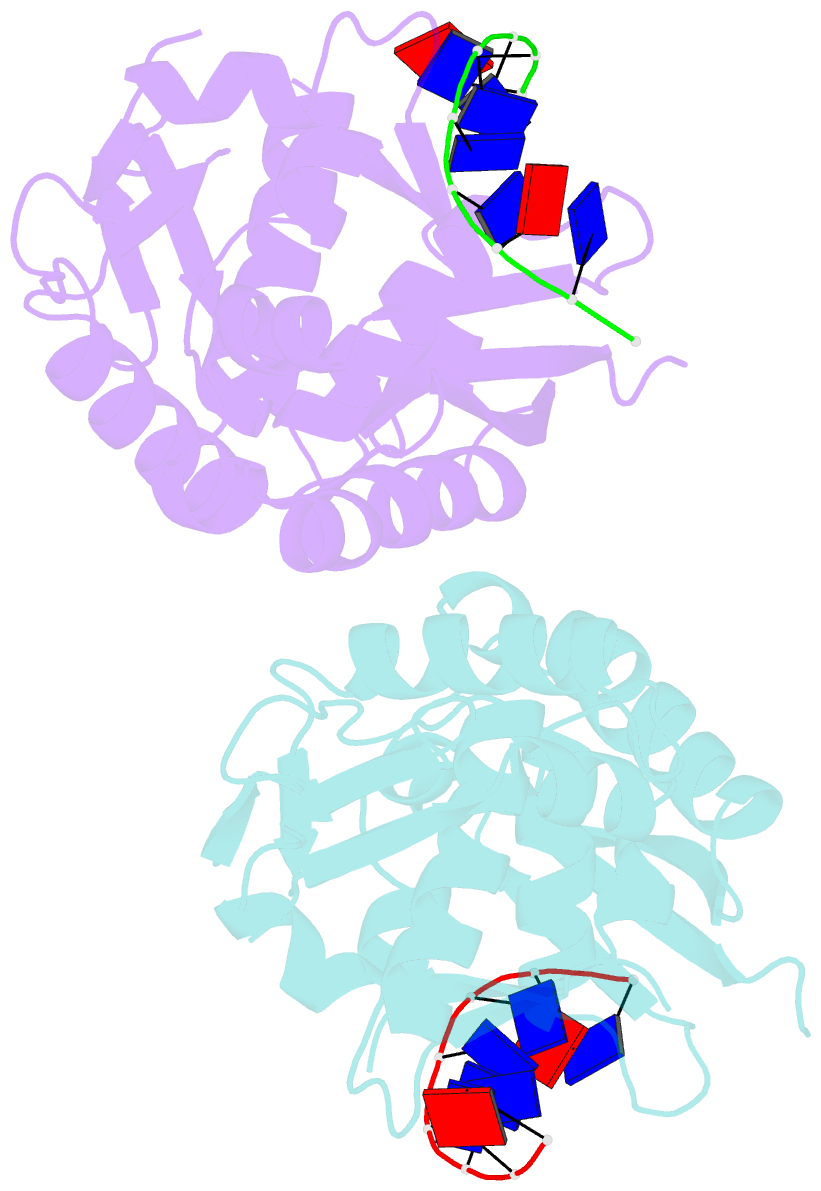
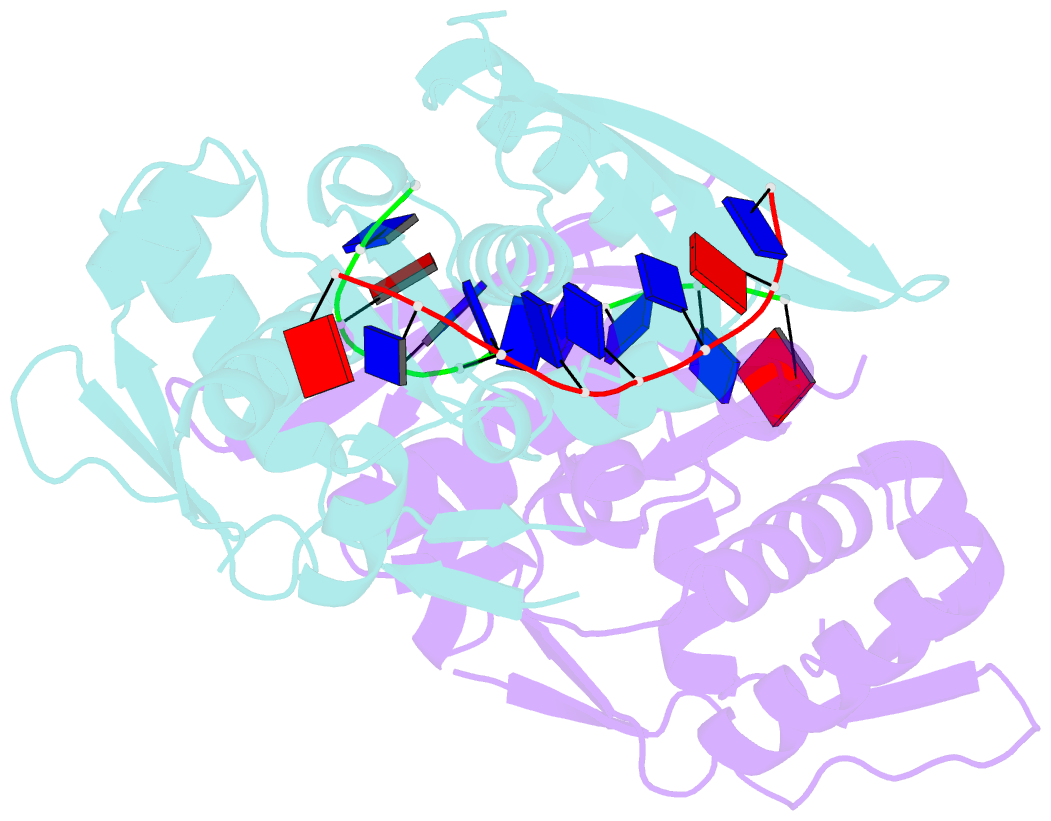
- The nucleoprotein complex of rep protein with due ssDNA
- Reference
- Wegrzyn K, Oliwa M, Nowacka M, Zabrocka E, Bury K, Purzycki P, Czaplewska P, Pipka J, Giraldo R, Konieczny I (2023): "Rep protein accommodates together dsDNA and ssDNA which enables a loop-back mechanism to plasmid DNA replication initiation." Nucleic Acids Res., 51, 10551-10567. doi: 10.1093/nar/gkad740.
- Abstract
- For DNA replication initiation in Bacteria, replication initiation proteins bind to double-stranded DNA (dsDNA) and interact with single-stranded DNA (ssDNA) at the replication origin. The structural-functional relationship of the nucleoprotein complex involving initiator proteins is still elusive and different models are proposed. In this work, based on crosslinking combined with mass spectrometry (MS), the analysis of mutant proteins and crystal structures, we defined amino acid residues essential for the interaction between plasmid Rep proteins, TrfA and RepE, and ssDNA. This interaction and Rep binding to dsDNA could not be provided in trans, and both are important for dsDNA melting at DNA unwinding element (DUE). We solved two crystal structures of RepE: one in a complex with ssDNA DUE, and another with both ssDNA DUE and dsDNA containing RepE-specific binding sites (iterons). The amino acid residues involved in interaction with ssDNA are located in the WH1 domain in stand β1, helices α1 and α2 and in the WH2 domain in loops preceding strands β1' and β2' and in these strands. It is on the opposite side compared to RepE dsDNA-recognition interface. Our data provide evidence for a loop-back mechanism through which the plasmid replication initiator molecule accommodates together dsDNA and ssDNA.