Summary information and primary citation
- PDB-id
-
8c00;
DSSR-derived features in text and
JSON formats; DNAproDB
- Class
- ribosome
- Method
- cryo-EM (2.9 Å)
- Summary
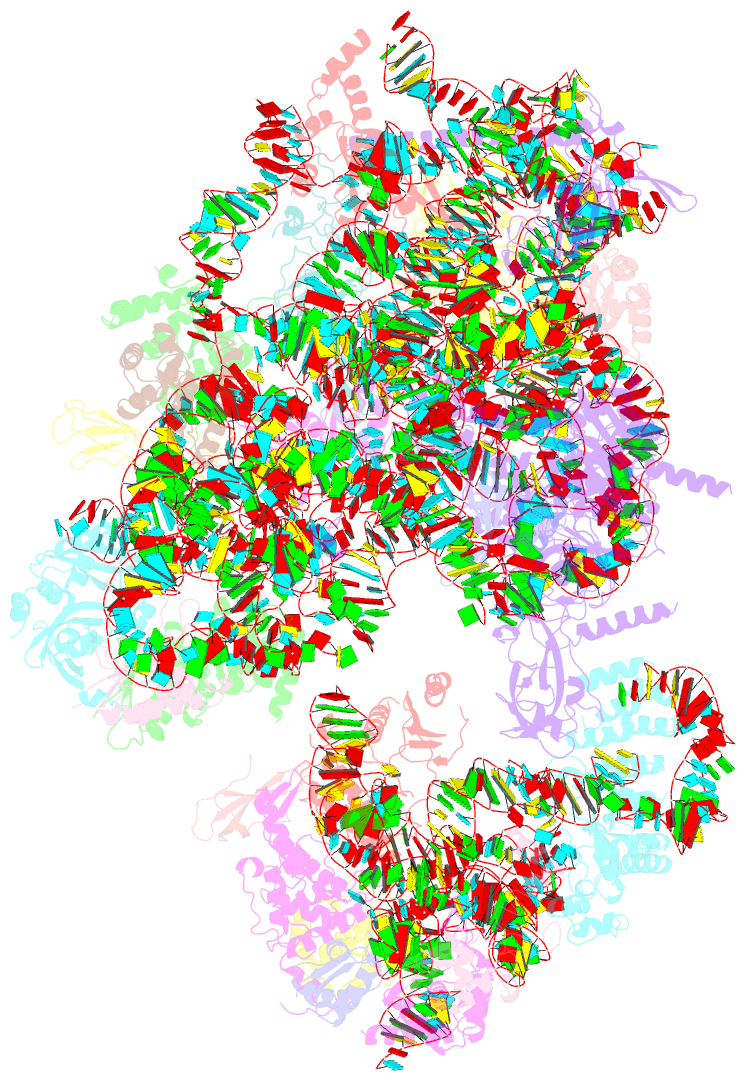
- Enp1tap-s21_a population of yeast small ribosomal
subunit precursors depleted of rps21-es21
- Reference
-
Poll G, Griesenbeck J, Tschochner H, Milkereit P (2023):
"Impact of
the yeast S0/uS2-cluster ribosomal protein rpS21/eS21 on
rRNA folding and the architecture of small ribosomal
subunit precursors." Plos One,
18, e0283698. doi: 10.1371/journal.pone.0283698.
- Abstract
- RpS0/uS2, rpS2/uS5, and rpS21/eS21 form a cluster of
ribosomal proteins (S0-cluster) at the head-body junction
near the central pseudoknot of eukaryotic small ribosomal
subunits (SSU). Previous work in yeast indicated that
S0-cluster assembly is required for the stabilisation and
maturation of SSU precursors at specific post-nucleolar
stages. Here, we analysed the role of S0-cluster formation
for rRNA folding. Structures of SSU precursors isolated
from yeast S0-cluster expression mutants or control strains
were analysed by cryogenic electron microscopy. The
obtained resolution was sufficient to detect individual
2'-O-methyl RNA modifications using an unbiased scoring
approach. The data show how S0-cluster formation enables
the initial recruitment of the pre-rRNA processing factor
Nob1 in yeast. Furthermore, they reveal hierarchical
effects on the pre-rRNA folding pathway, including the
final maturation of the central pseudoknot. Based on these
structural insights we discuss how formation of the
S0-cluster determines at this early cytoplasmic assembly
checkpoint if SSU precursors further mature or are
degraded.