Summary information and primary citation
- PDB-id
- 8e95; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- transcription, transferase-DNA-RNA
- Method
- cryo-EM (2.9 Å)
- Summary
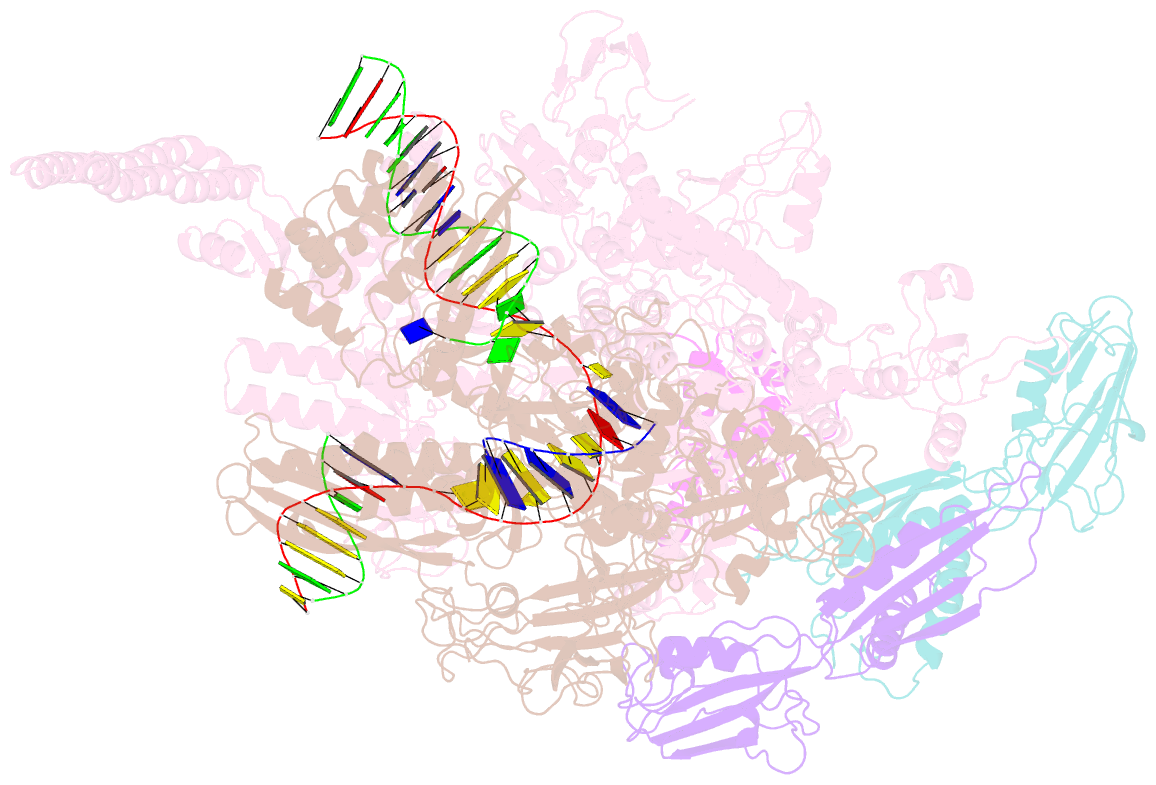
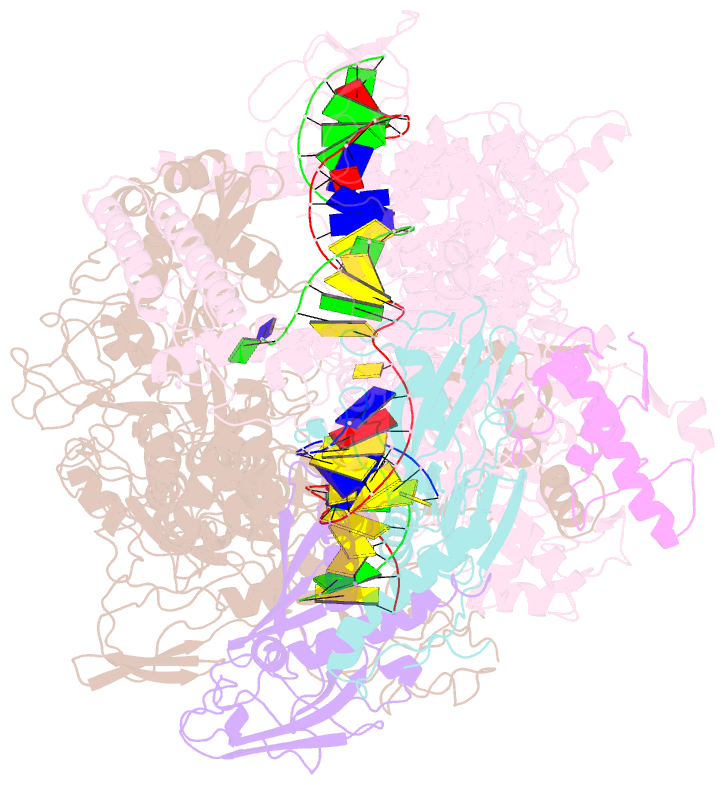
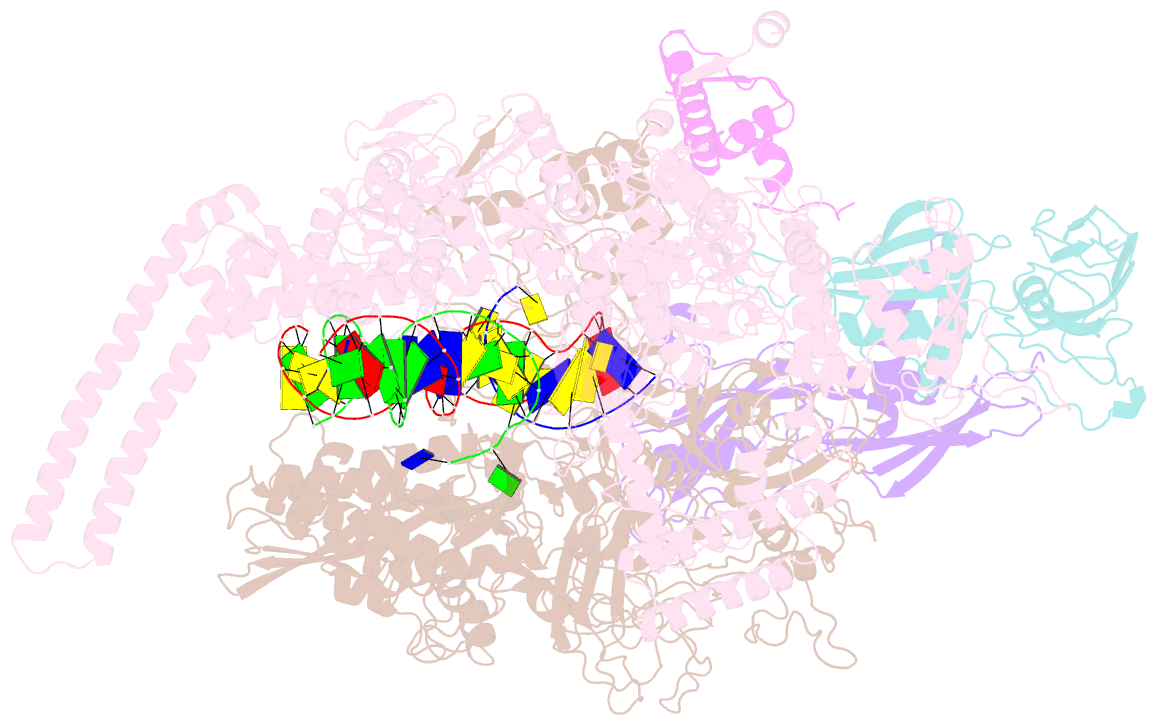
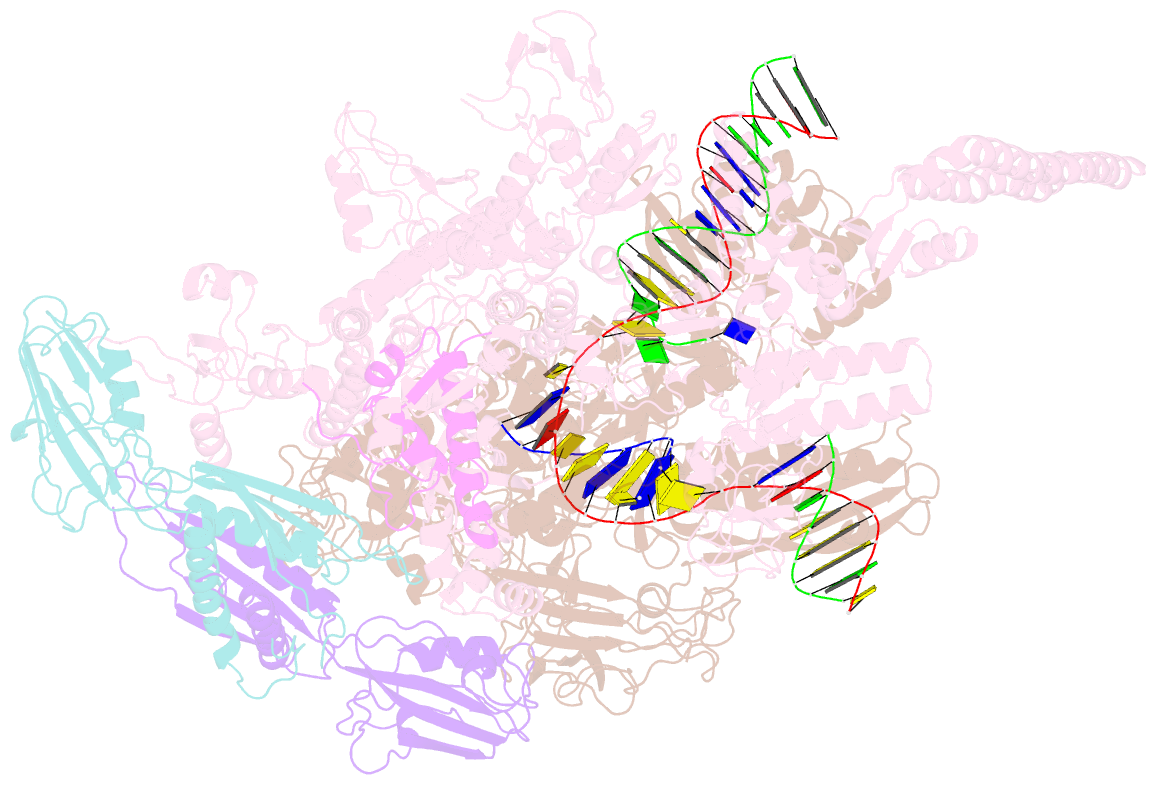
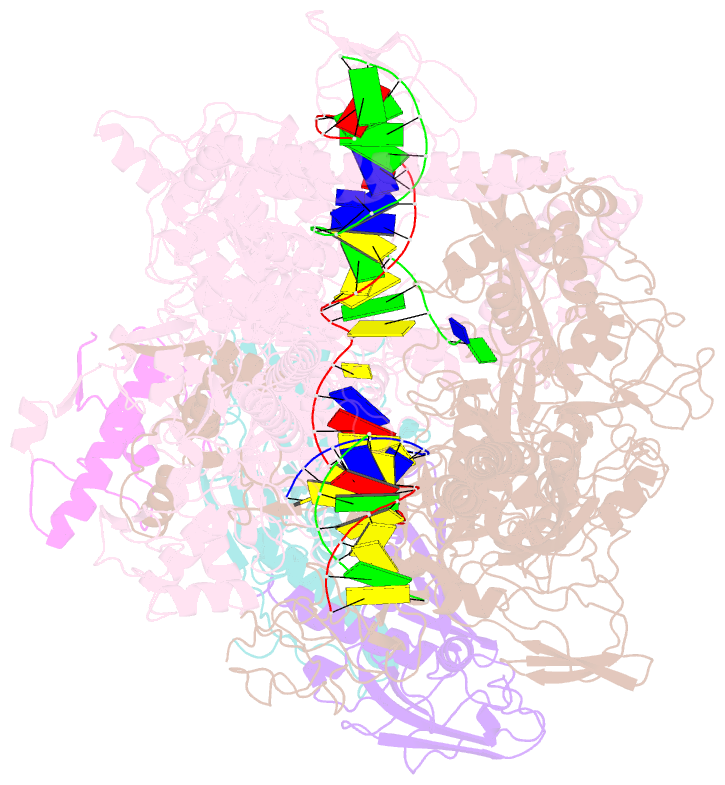
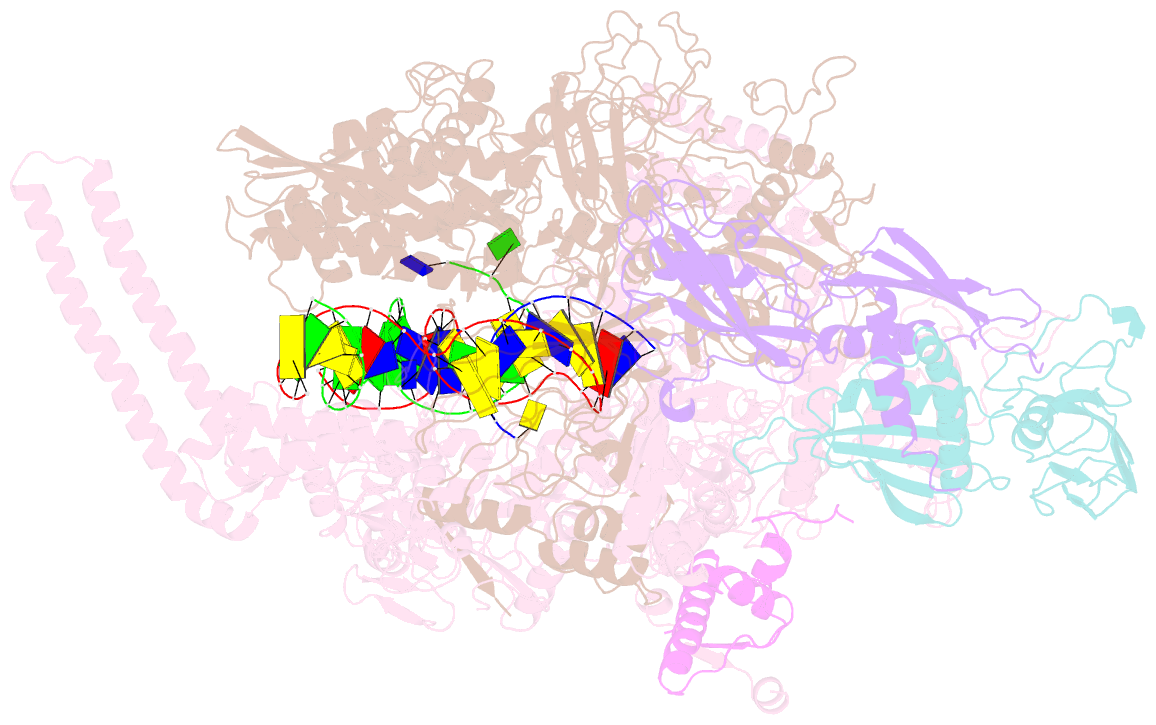
- Mycobacterium tuberculosis rnap elongation complex
- Reference
- Delbeau M, Omollo EO, Froom R, Koh S, Mooney RA, Lilic M, Brewer JJ, Rock J, Darst SA, Campbell EA, Landick R (2023): "Structural and functional basis of the universal transcription factor NusG pro-pausing activity in Mycobacterium tuberculosis." Mol.Cell, 83, 1474-1488.e8. doi: 10.1016/j.molcel.2023.04.007.
- Abstract
- Transcriptional pauses mediate regulation of RNA biogenesis. DNA-encoded pause signals trigger pausing by stabilizing RNA polymerase (RNAP) swiveling and inhibiting DNA translocation. The N-terminal domain (NGN) of the only universal transcription factor, NusG/Spt5, modulates pausing through contacts to RNAP and DNA. Pro-pausing NusGs enhance pauses, whereas anti-pausing NusGs suppress pauses. Little is known about pausing and NusG in the human pathogen Mycobacterium tuberculosis (Mtb). We report that MtbNusG is pro-pausing. MtbNusG captures paused, swiveled RNAP by contacts to the RNAP protrusion and nontemplate-DNA wedged between the NGN and RNAP gate loop. In contrast, anti-pausing Escherichia coli (Eco) NGN contacts the MtbRNAP gate loop, inhibiting swiveling and pausing. Using CRISPR-mediated genetics, we show that pro-pausing NGN is required for mycobacterial fitness. Our results define an essential function of mycobacterial NusG and the structural basis of pro- versus anti-pausing NusG activity, with broad implications for the function of all NusG orthologs.