Summary information and primary citation
- PDB-id
- 8h40; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- transcription
- Method
- cryo-EM (3.6 Å)
- Summary
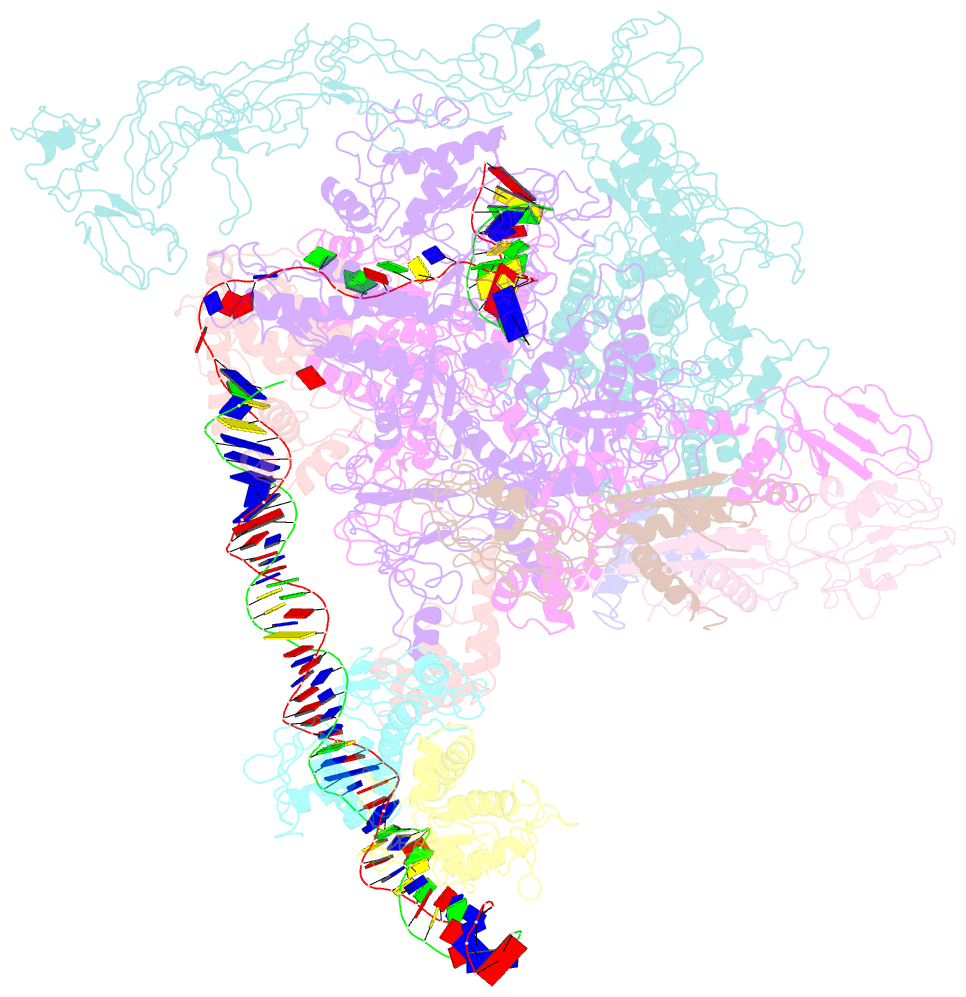
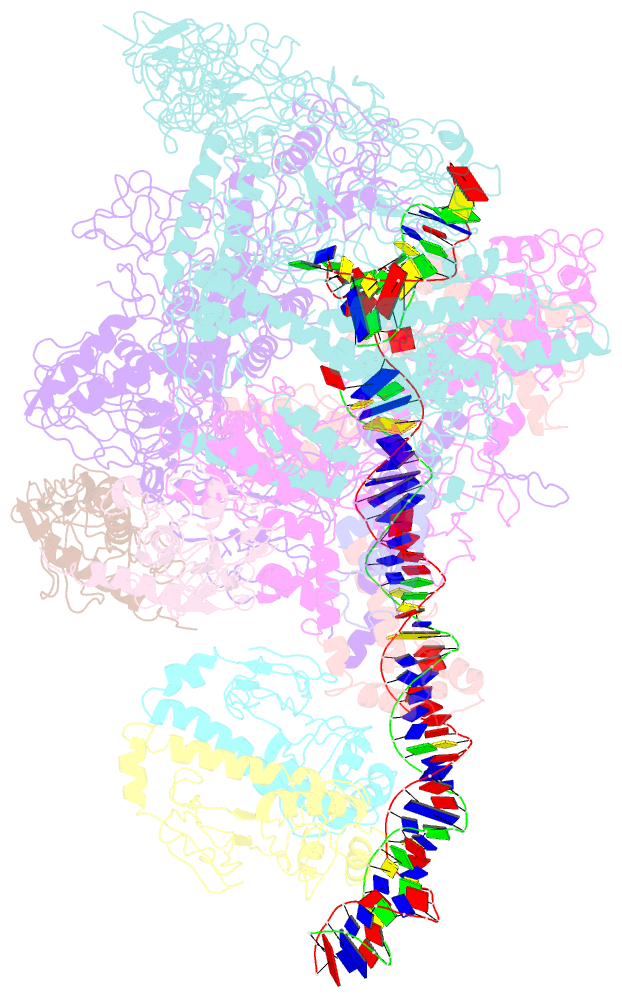
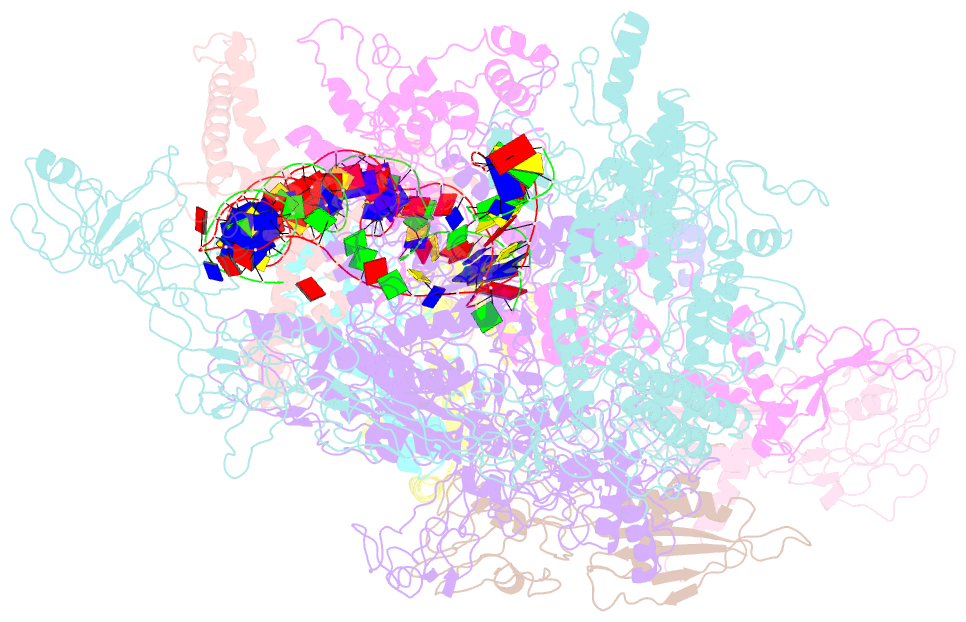
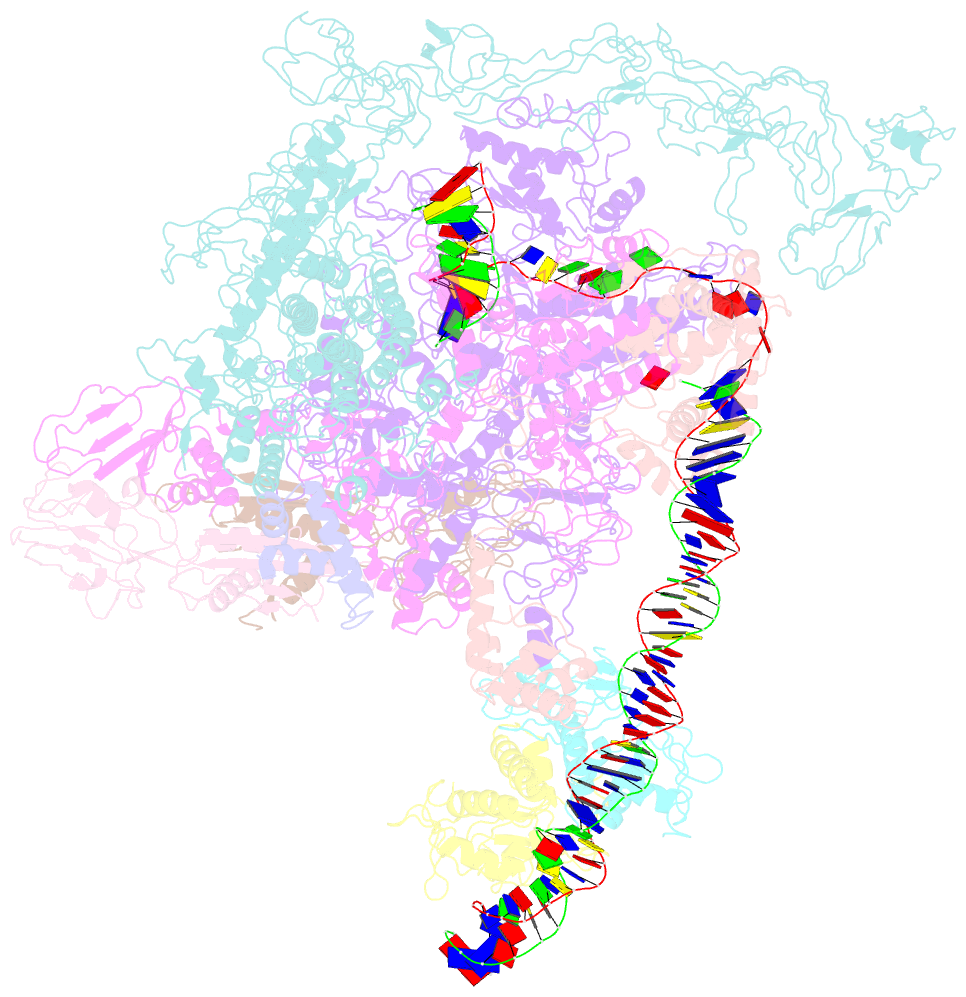
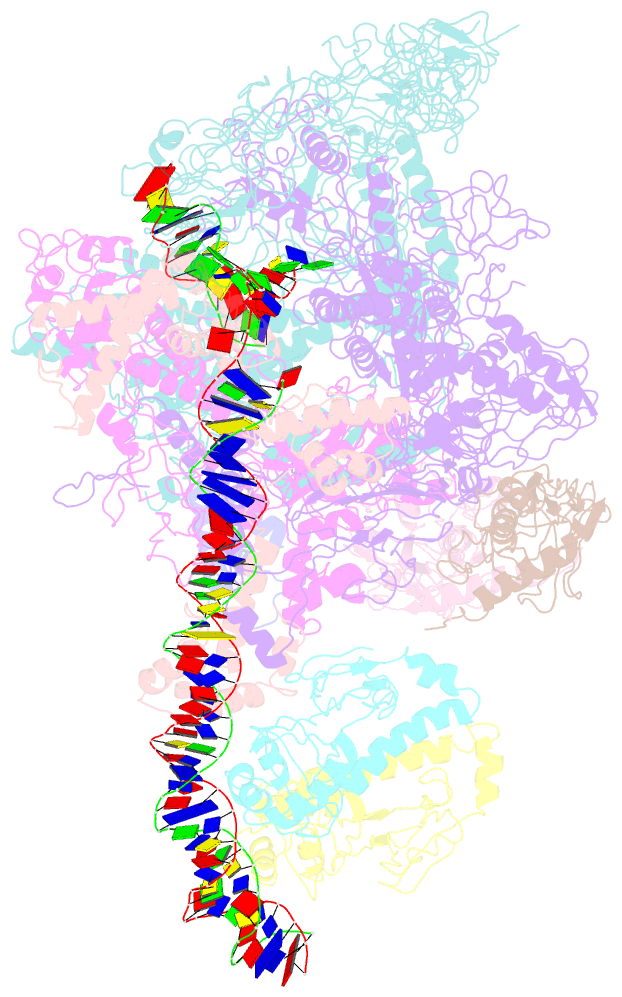
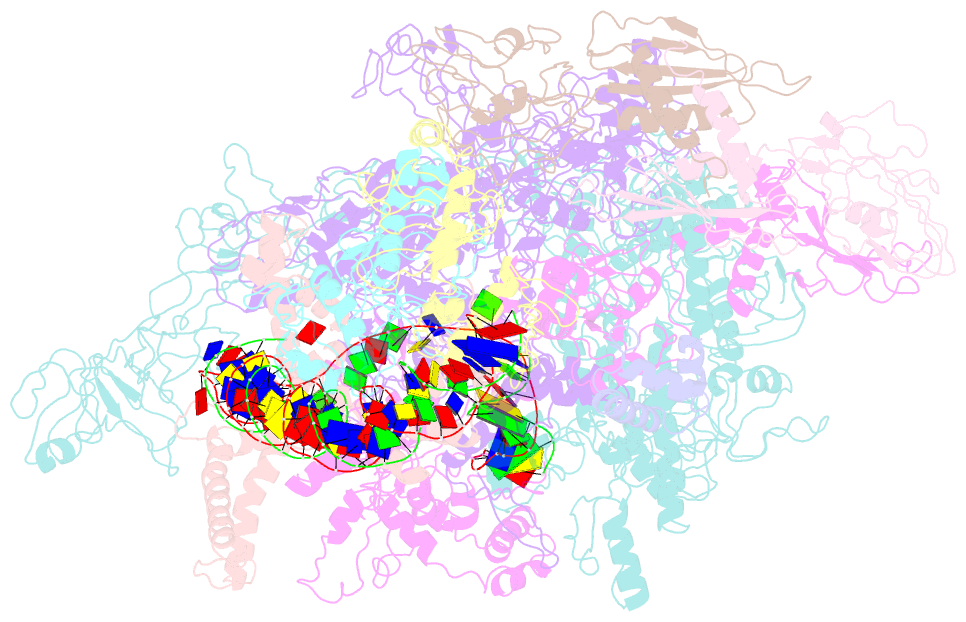
- cryo-EM structure of the transcription activation complex ntca-tac
- Reference
- Han SJ, Jiang YL, You LL, Shen LQ, Wu X, Yang F, Cui N, Kong WW, Sun H, Zhou K, Meng HC, Chen ZP, Chen Y, Zhang Y, Zhou CZ (2024): "DNA looping mediates cooperative transcription activation." Nat.Struct.Mol.Biol., 31, 293-299. doi: 10.1038/s41594-023-01149-7.
- Abstract
- Transcription factors respond to multilevel stimuli and co-occupy promoter regions of target genes to activate RNA polymerase (RNAP) in a cooperative manner. To decipher the molecular mechanism, here we report two cryo-electron microscopy structures of Anabaena transcription activation complexes (TACs): NtcA-TAC composed of RNAP holoenzyme, promoter and a global activator NtcA, and NtcA-NtcB-TAC comprising an extra context-specific regulator, NtcB. Structural analysis showed that NtcA binding makes the promoter DNA bend by ∼50°, which facilitates RNAP to contact NtcB at the distal upstream NtcB box. The sequential binding of NtcA and NtcB induces looping back of promoter DNA towards RNAP, enabling the assembly of a fully activated TAC bound with two activators. Together with biochemical assays, we propose a 'DNA looping' mechanism of cooperative transcription activation in bacteria.