Summary information and primary citation
- PDB-id
- 8k60; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- gene regulation
- Method
- cryo-EM (3.4 Å)
- Summary
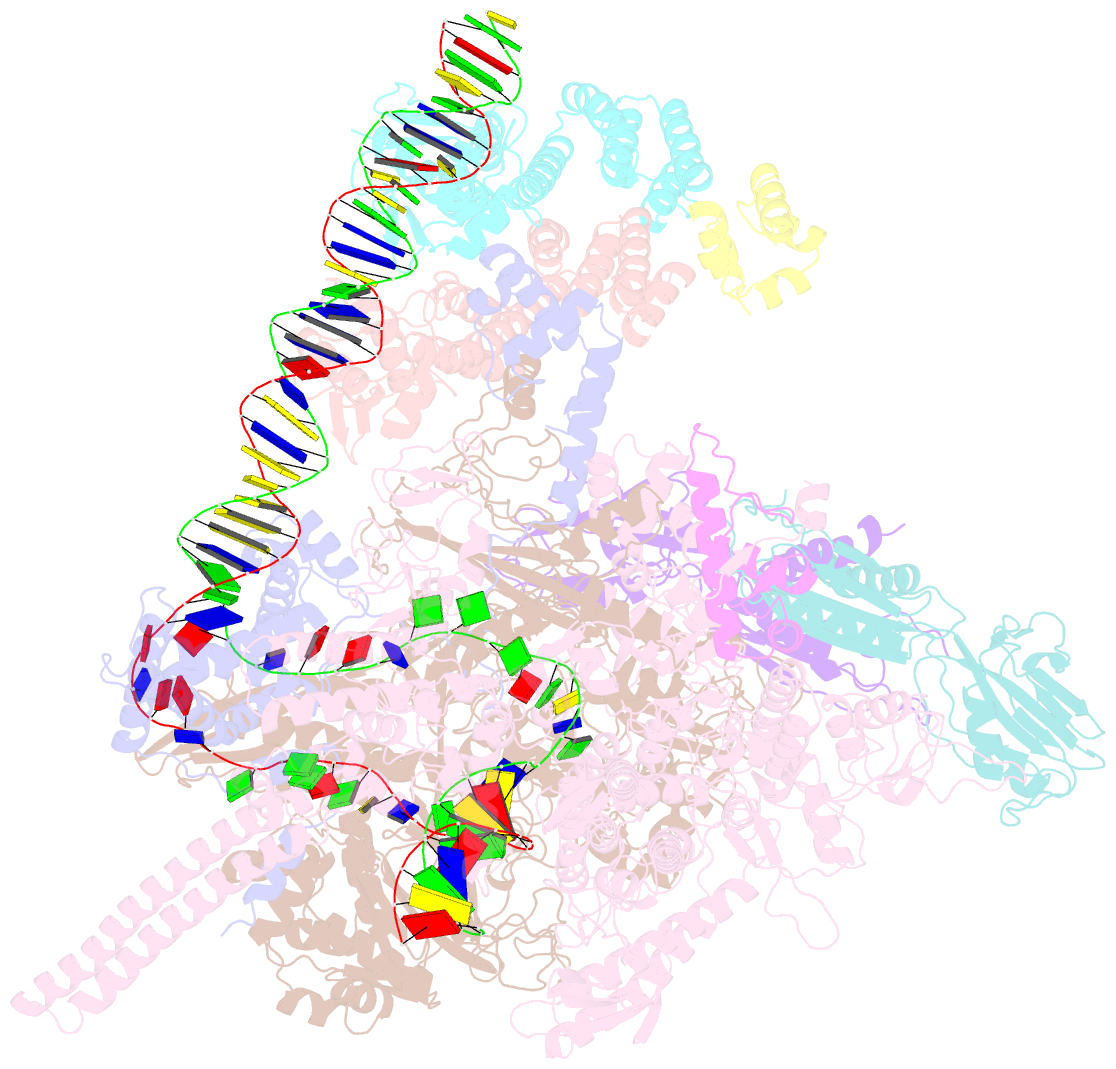
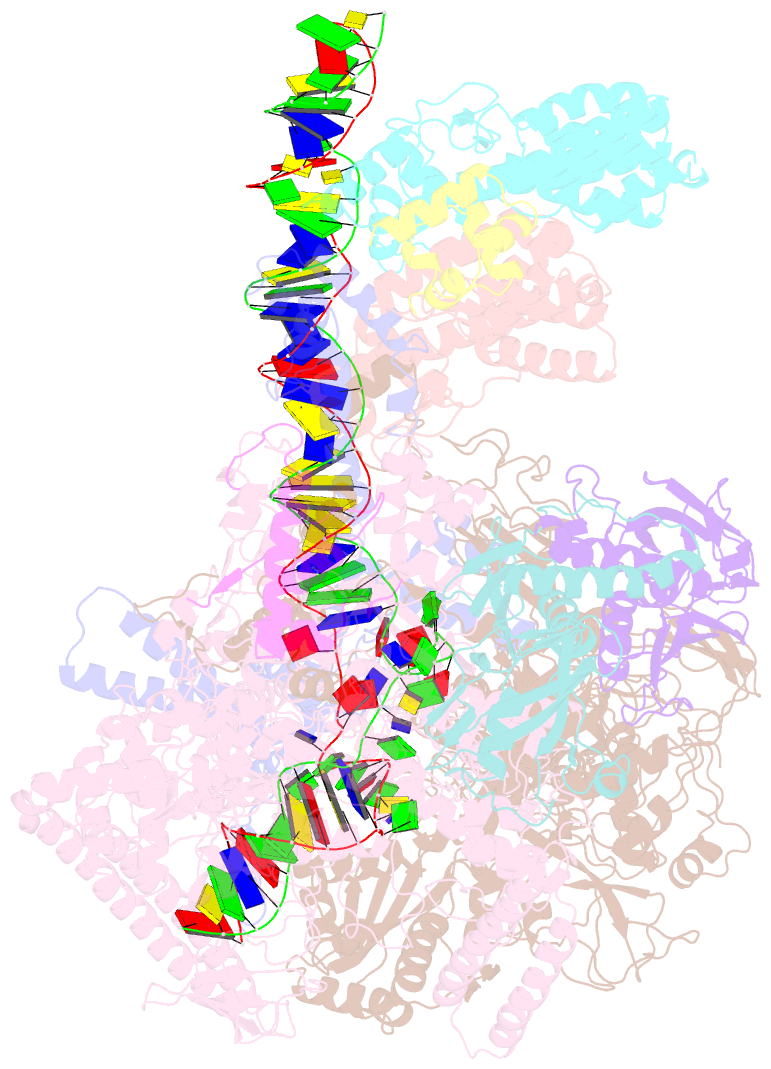
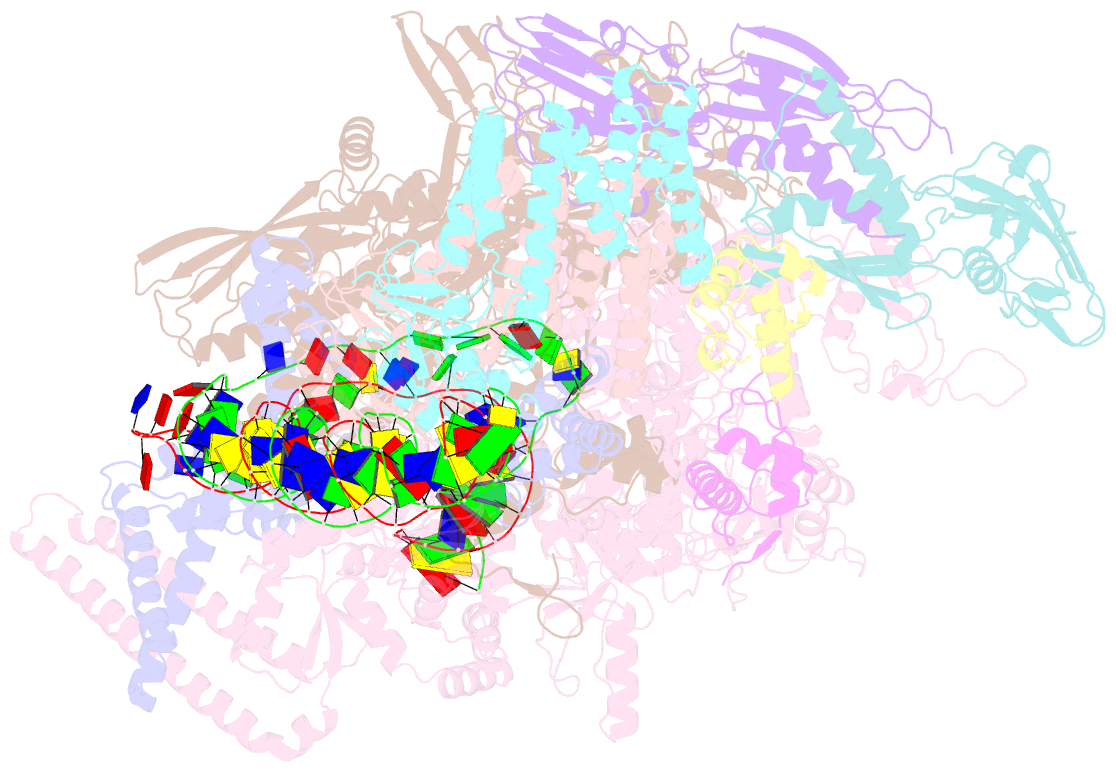
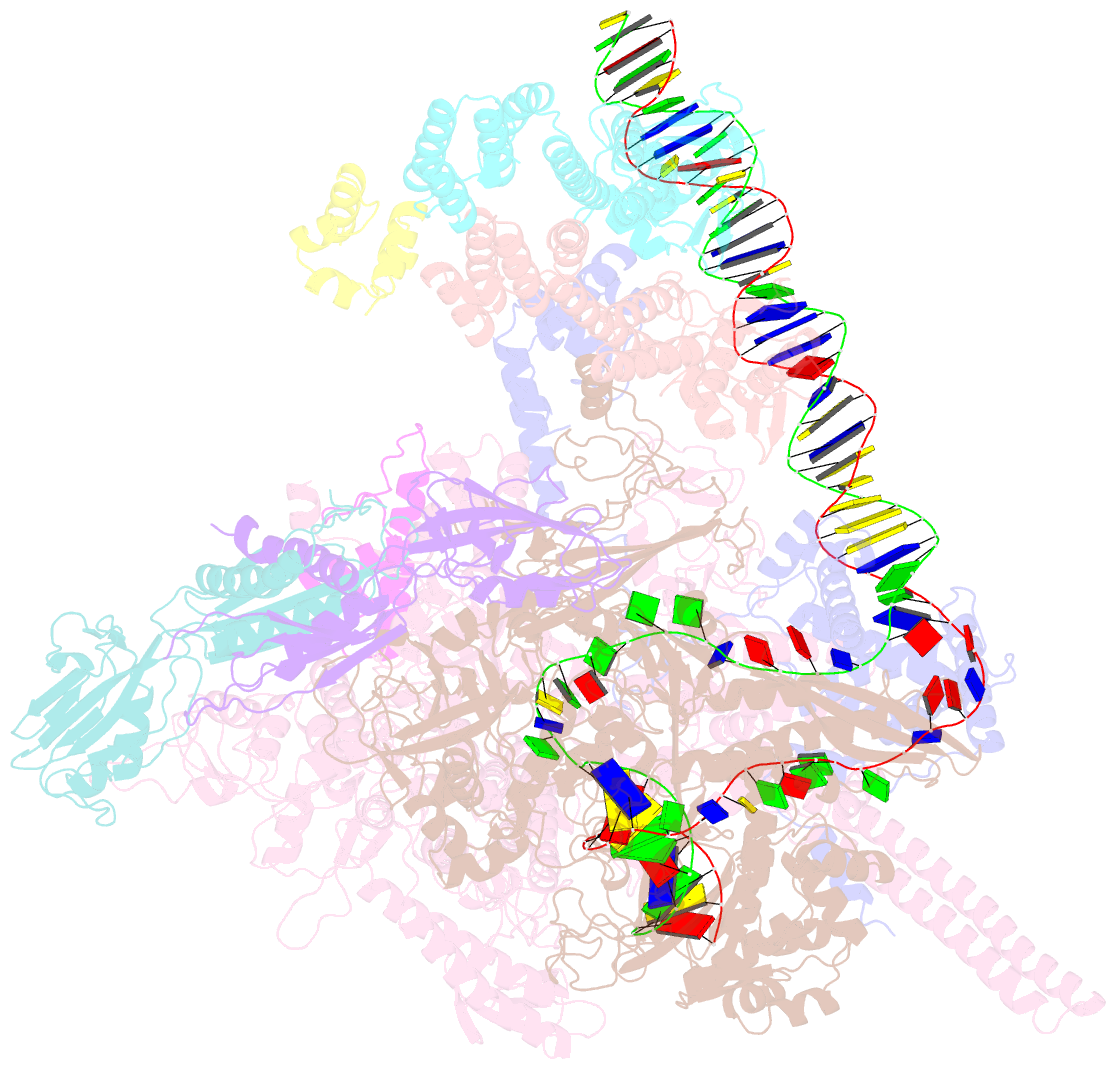
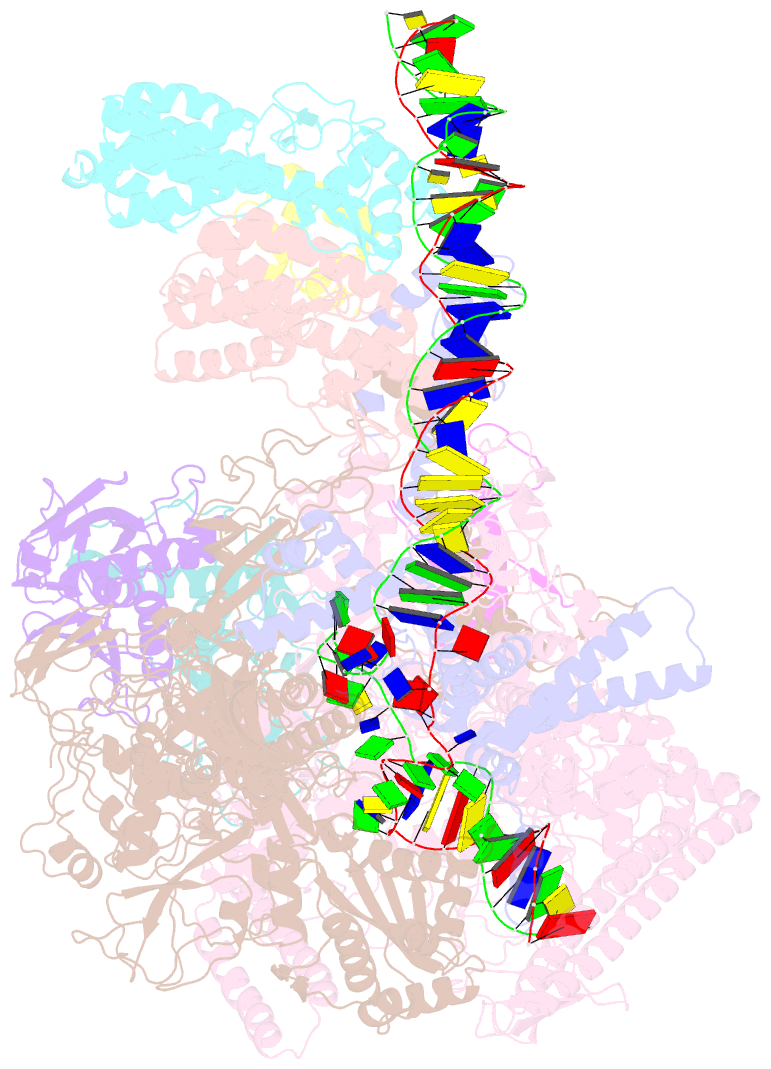
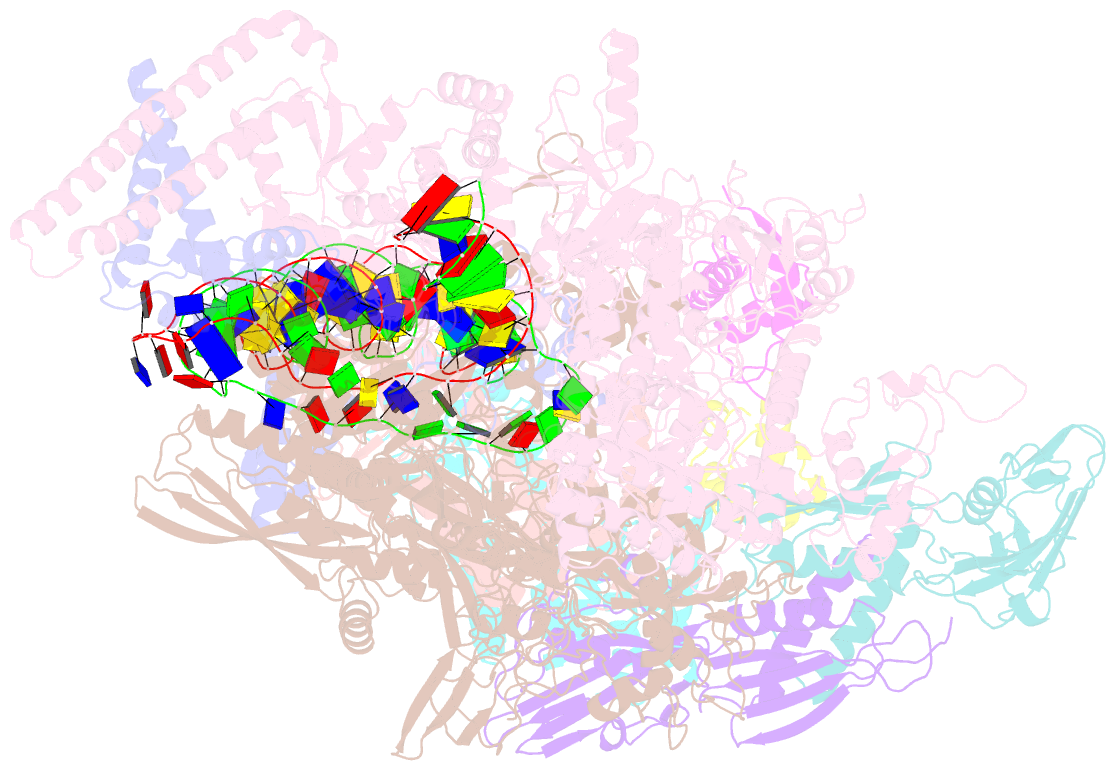
- cryo-EM structure of streptomyces coelicolor transcription initiation complex with the global transcription factor afsr
- Reference
- Shi J, Ye Z, Feng Z, Wen A, Wang L, Zhang Z, Xu L, Song Q, Wang F, Liu T, Wang S, Feng Y, Lin W (2024): "Structural insights into transcription activation of the Streptomyces antibiotic regulatory protein, AfsR." Iscience, 27, 110421. doi: 10.1016/j.isci.2024.110421.
- Abstract
- The Streptomyces antibiotic regulatory proteins (SARPs) are ubiquitously distributed transcription activators in Streptomyces and control antibiotics biosynthesis and morphological differentiation. However, the molecular mechanism behind SARP-dependent transcription initiation remains elusive. We here solve the cryo-EM structure of an AfsR-loading RNA polymerase (RNAP)-promoter intermediate complex (AfsR-RPi) including the Streptomyces coelicolor RNAP, a large SARP member AfsR, and its target promoter DNA that retains the upstream portion straight. The structure reveals that one dimeric N-terminal AfsR-SARP domain (AfsR-SARP) specifically engages with the same face of the AfsR-binding sites by the conserved DNA-binding domains (DBDs), replacing σHrdBR4 to bind the suboptimal -35 element, and shortens the spacer between the -10 and -35 elements. Notably, the AfsR-SARPs also recruit RNAP through extensively interacting with its conserved domains (β flap, σHrdBR4, and αCTD). Thus, these macromolecular snapshots support a general model and provide valuable clues for SARP-dependent transcription activation in Streptomyces.