Summary information and primary citation
- PDB-id
- 8mht; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- transferase-DNA
- Method
- X-ray (2.76 Å)
- Summary
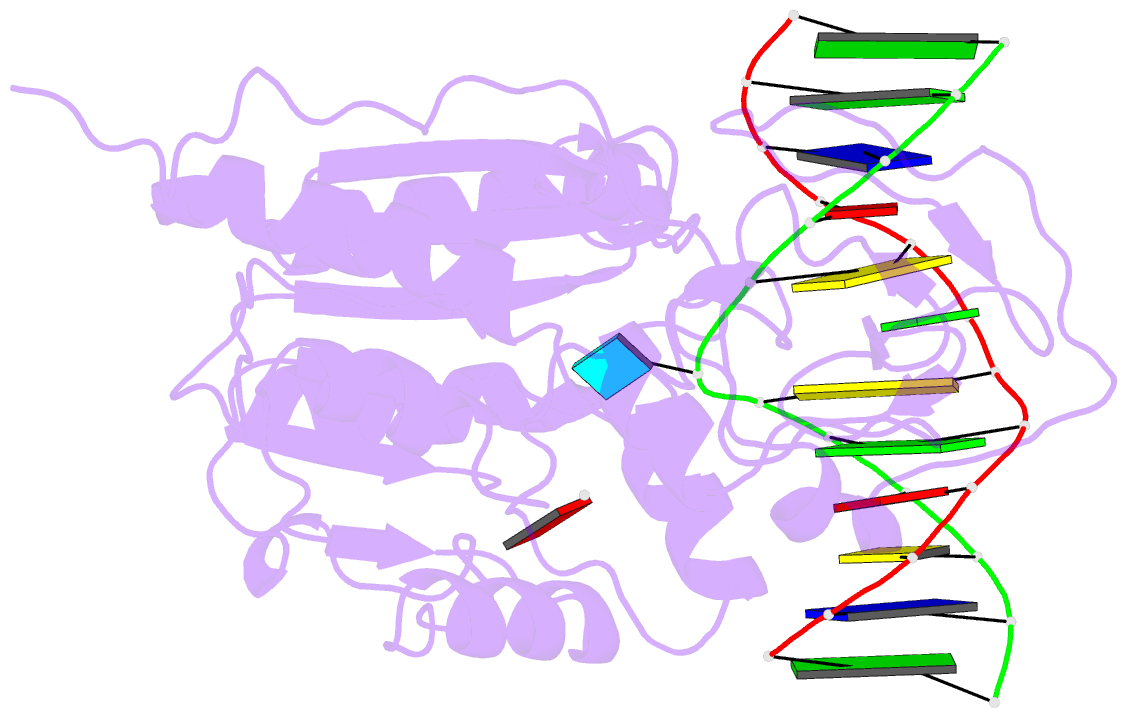
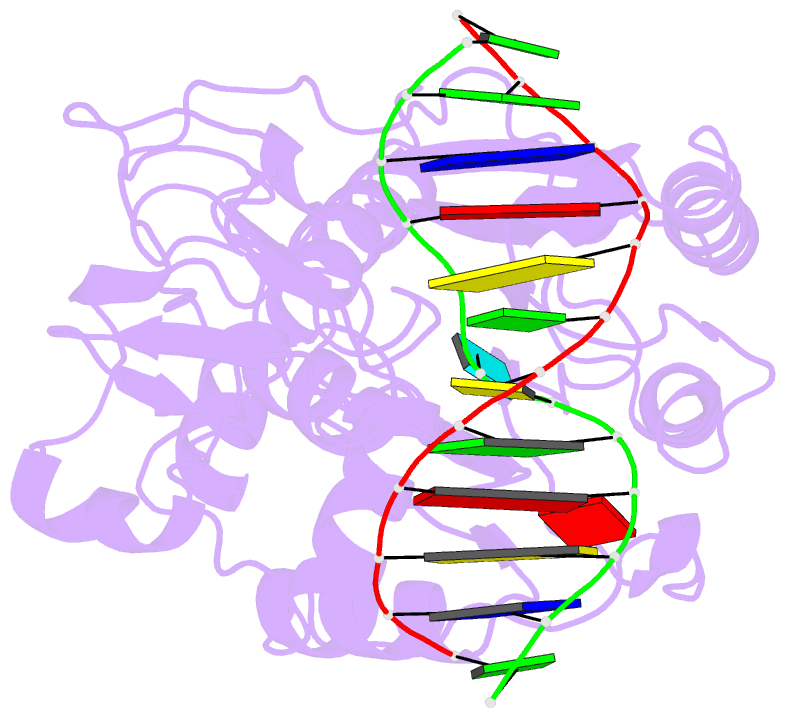
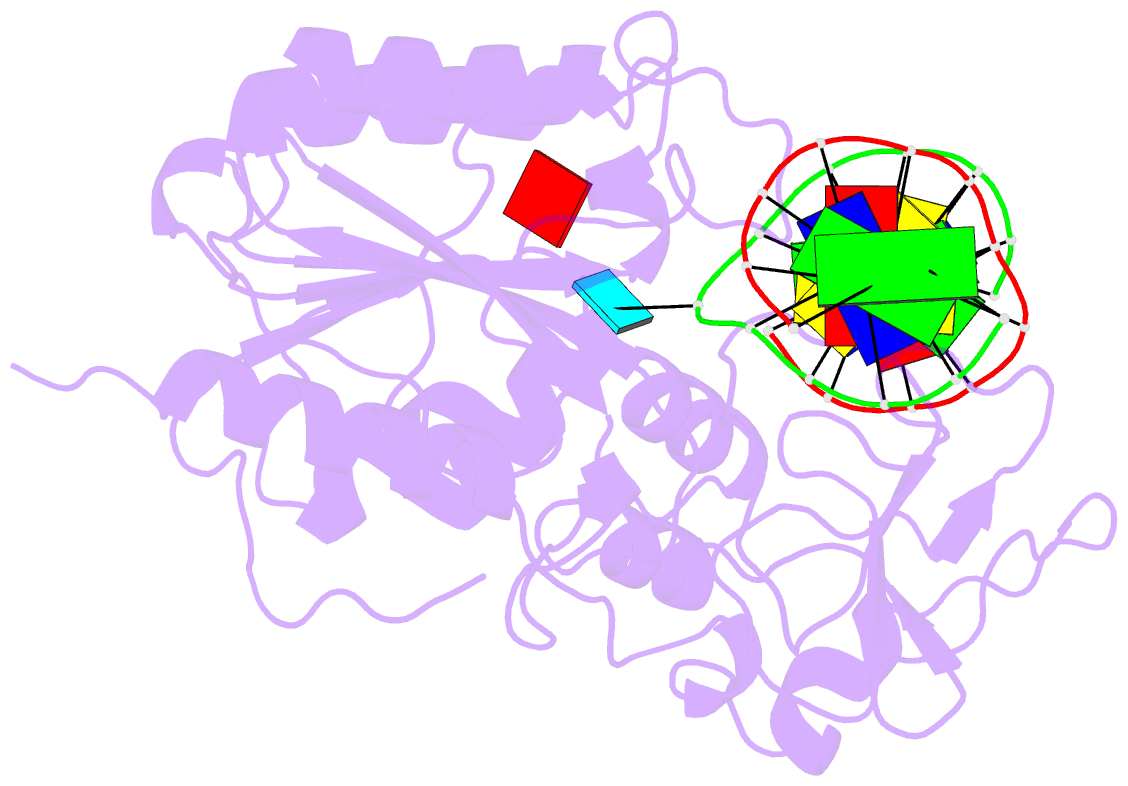
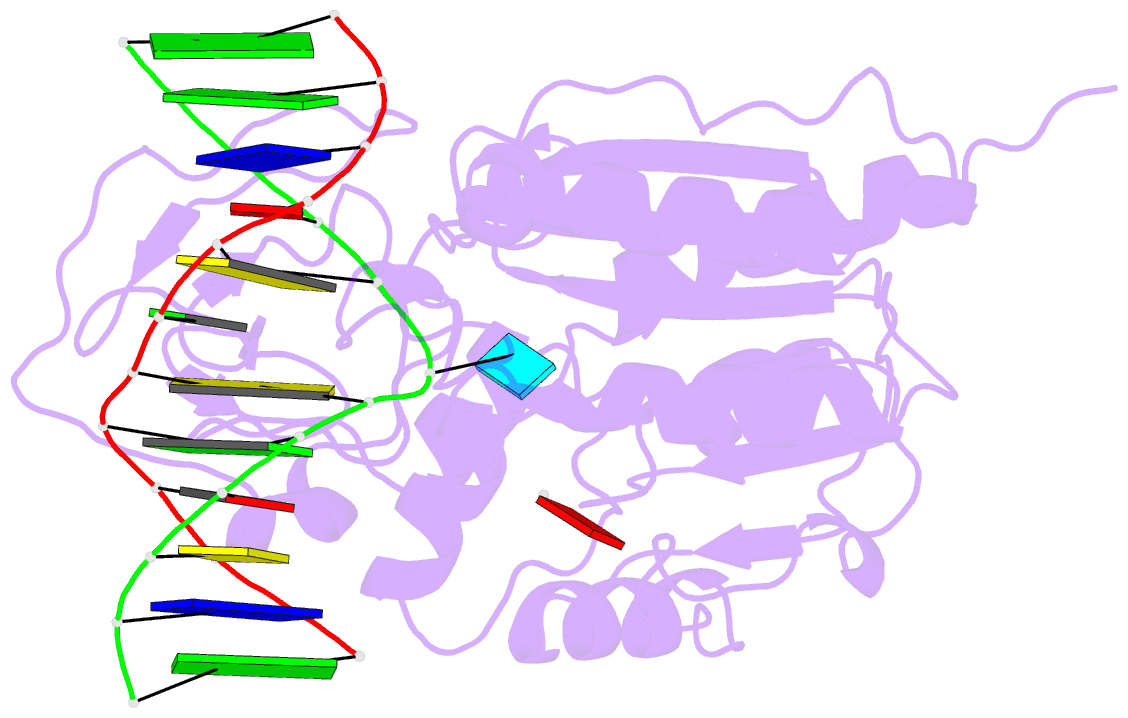
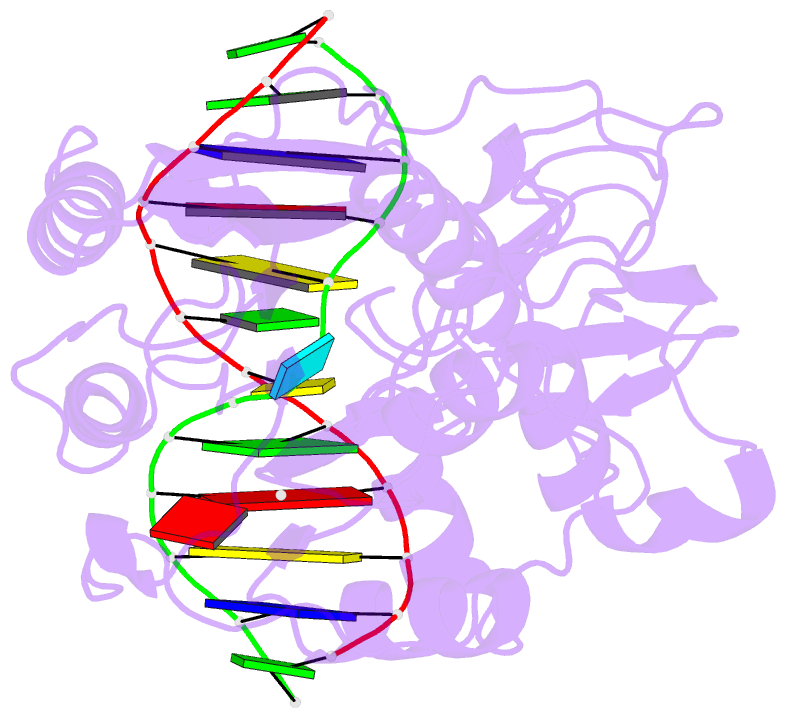
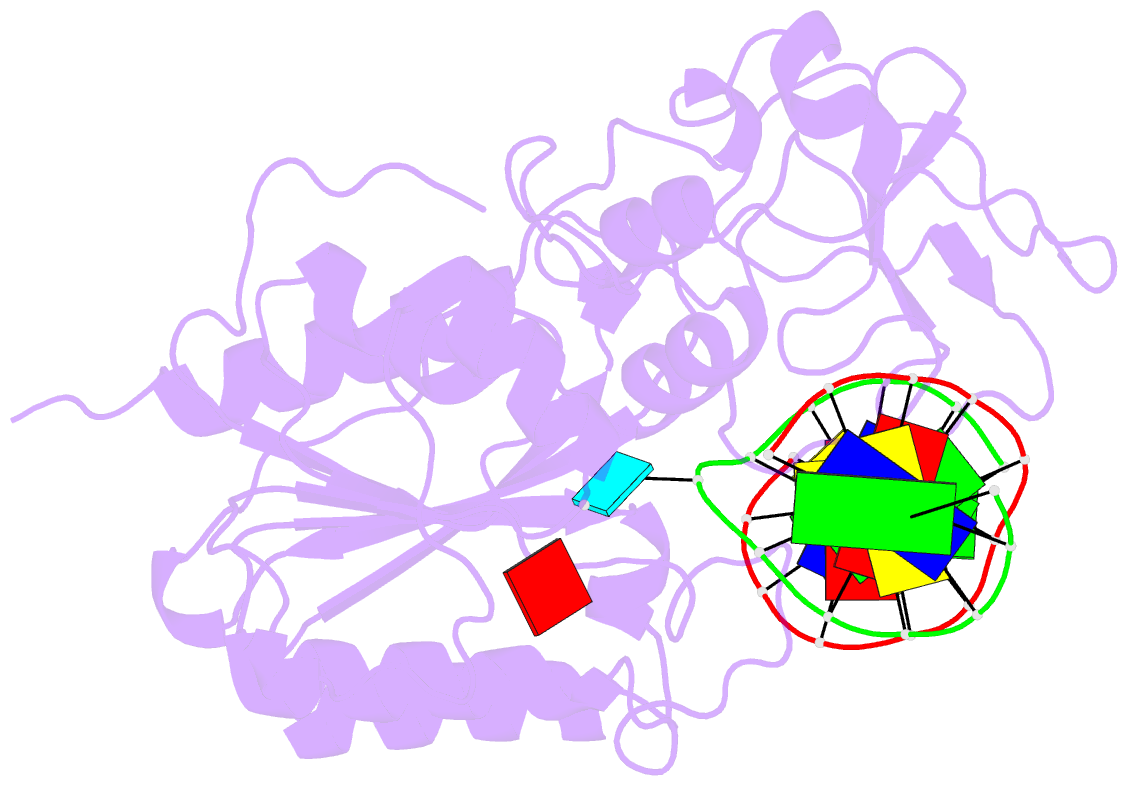
- Cytosine-specific methyltransferase hhai-DNA complex
- Reference
- O'Gara M, Horton JR, Roberts RJ, Cheng X (1998): "Structures of HhaI methyltransferase complexed with substrates containing mismatches at the target base." Nat.Struct.Biol., 5, 872-877. doi: 10.1038/2312.
- Abstract
- Three structures have been determined for complexes between HhaI methyltransferase (M.HhaI) and oligonucleotides containing a G:A, G:U or G:AP (AP = abasic or apurinic/apyrimidinic) mismatch at the target base pair. The mismatched adenine, uracil and abasic site are all flipped out of the DNA helix and located in the enzyme's active-site pocket, adopting the same conformation as in the flipped-out normal substrate. These results, particularly the flipped-out abasic deoxyribose sugar, provide insight into the mechanism of base flipping. If the process involves the protein pushing the base out of the helix, then the push must take place not on the base, but rather on the sugar-phosphate backbone. Thus rotation of the DNA backbone is probably the key to base flipping.