Summary information and primary citation
- PDB-id
-
8q3w;
DSSR-derived features in text and
JSON formats; DNAproDB
- Class
- DNA binding protein
- Method
- cryo-EM (3.18 Å)
- Summary
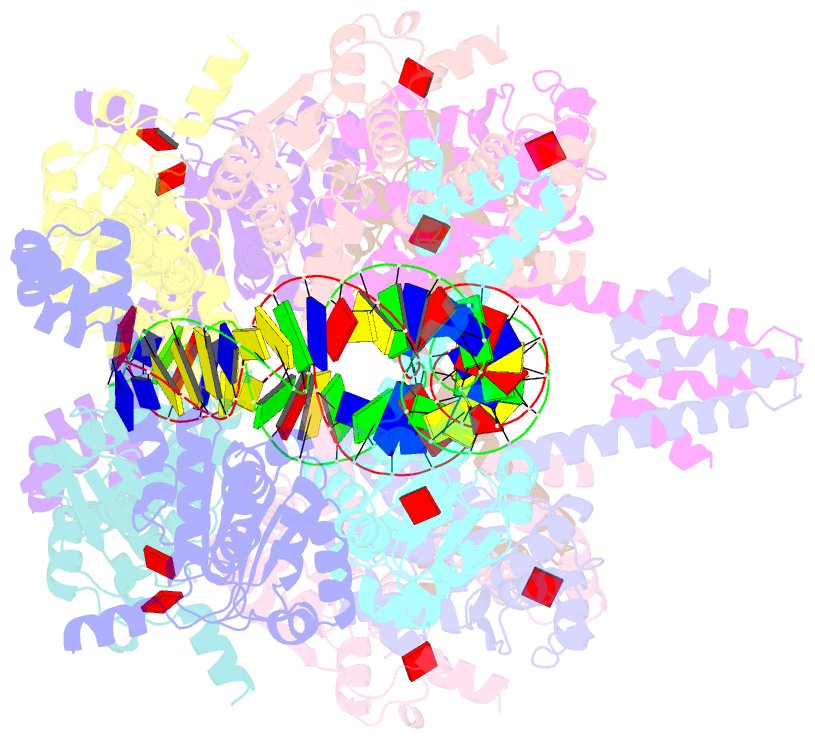
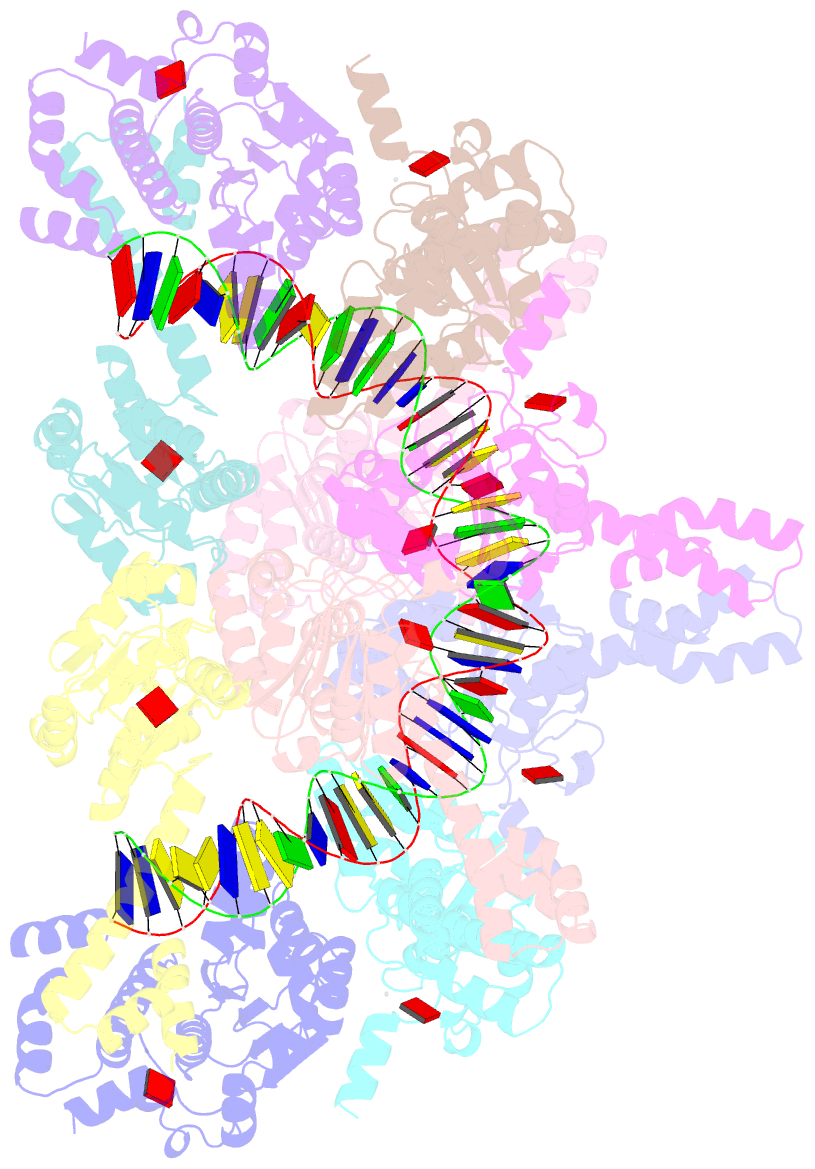
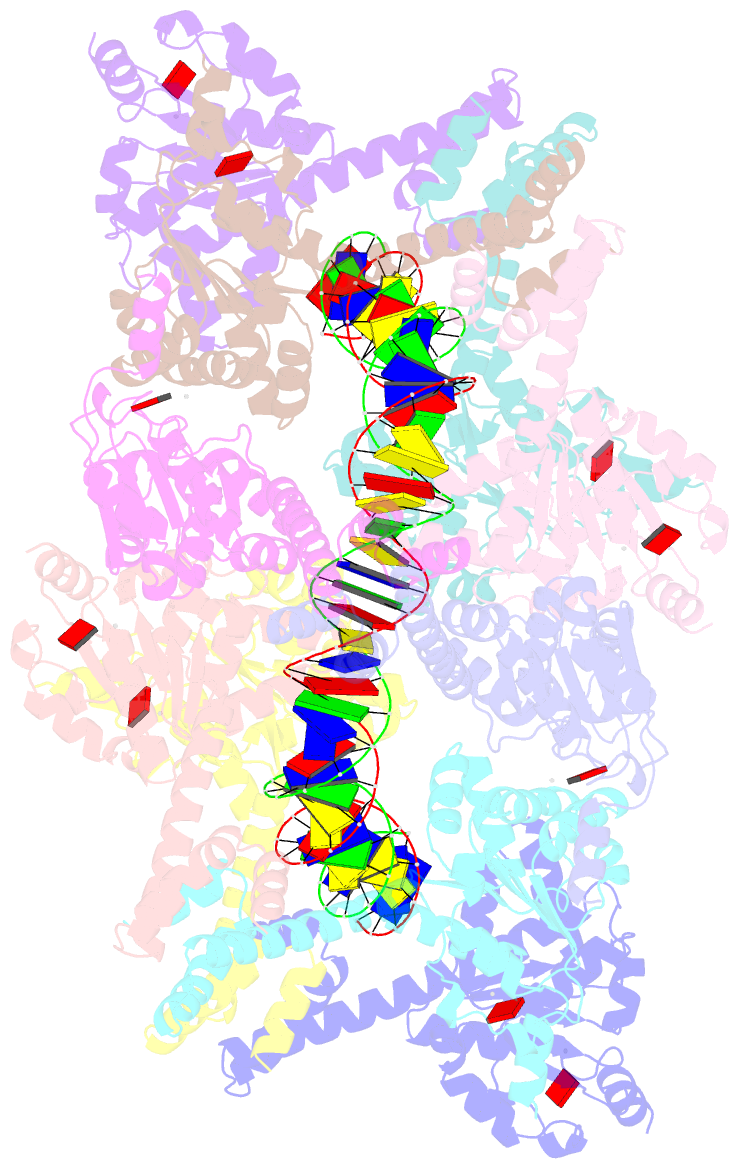
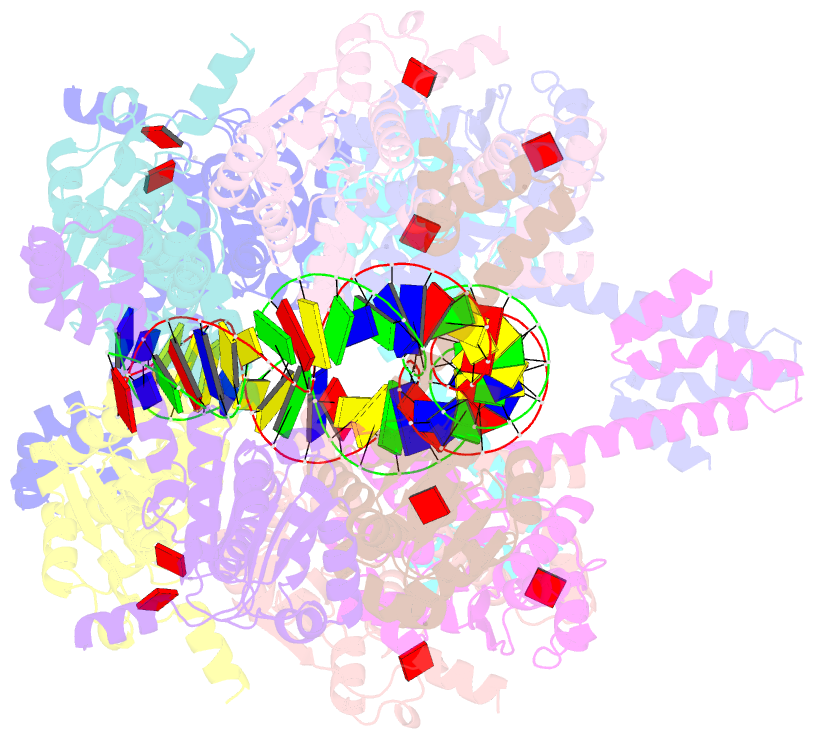
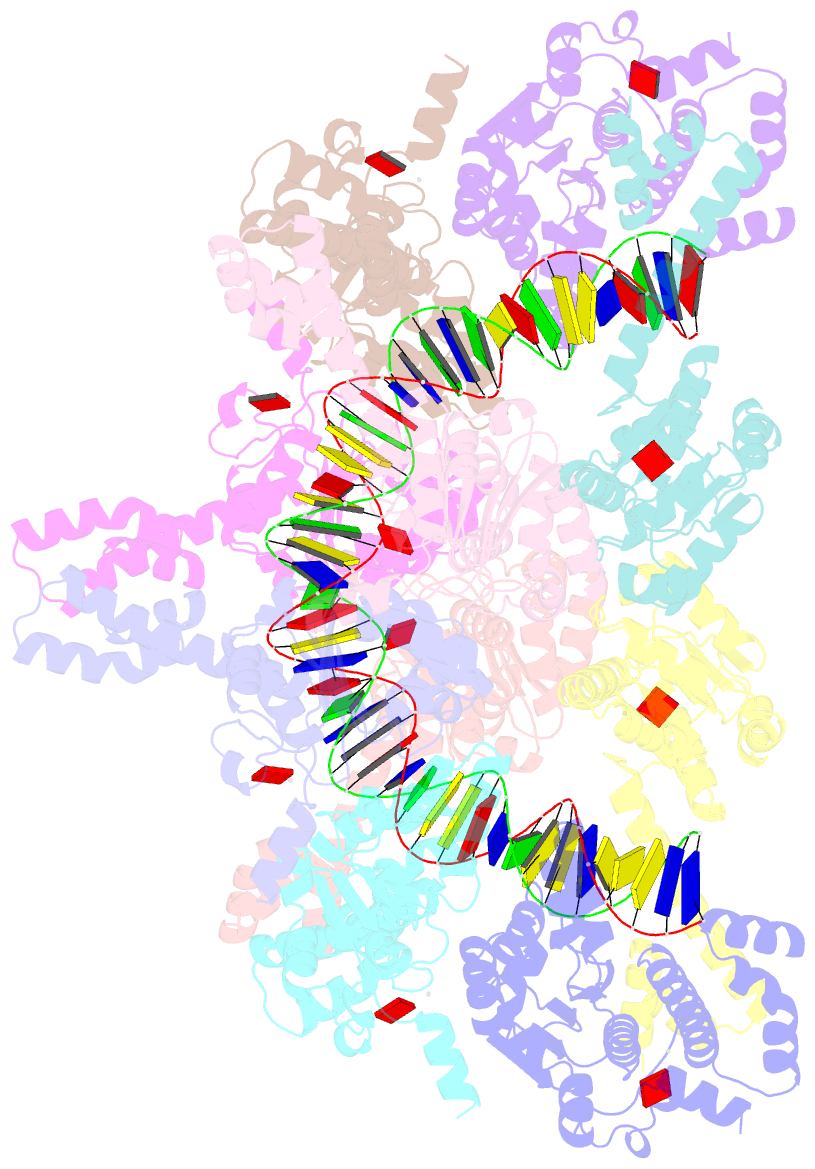
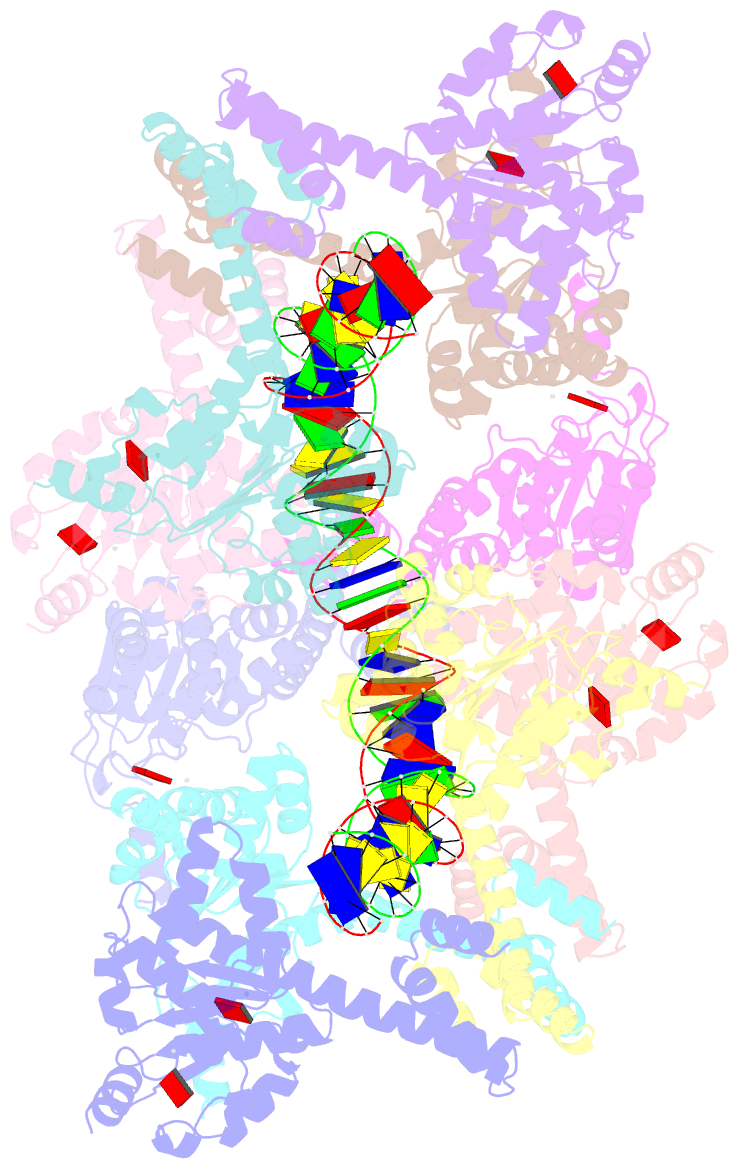
- Atp-bound istb in complex to duplex DNA
- Reference
-
de la Gandara A, Spinola-Amilibia M, Araujo-Bazan L,
Nunez-Ramirez R, Berger JM, Arias-Palomo E (2024):
"Molecular
basis for transposase activation by a dedicated AAA+
ATPase." Nature, 630,
1003-1011. doi: 10.1038/s41586-024-07550-6.
- Abstract
- Transposases drive chromosomal rearrangements and the
dissemination of drug-resistance genes and
toxins<sub>1-3</sub>. Although some
transposases act alone, many rely on dedicated AAA+ ATPase
subunits that regulate site selectivity and catalytic
function through poorly understood mechanisms. Using IS21
as a model transposase system, we show how an ATPase
regulator uses nucleotide-controlled assembly and DNA
deformation to enable structure-based site selectivity,
transposase recruitment, and activation and integration.
Solution and cryogenic electron microscopy studies show
that the IstB ATPase self-assembles into an
autoinhibited pentamer of dimers that tightly curves target
DNA into a half-coil. Two of these decamers dimerize, which
stabilizes the target nucleic acid into a kinked S-shaped
configuration that engages the IstA transposase at the
interface between the two IstB oligomers to form an
approximately 1 MDa transpososome complex. Specific
interactions stimulate regulator ATPase activity and
trigger a large conformational change on the transposase
that positions the catalytic site to perform DNA strand
transfer. These studies help explain how AAA+ ATPase
regulators-which are used by classical transposition
systems such as Tn7, Mu and CRISPR-associated elements-can
remodel their substrate DNA and cognate transposases to
promote function.