Summary information and primary citation
- PDB-id
- 8sxt; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- RNA binding protein-RNA-DNA
- Method
- cryo-EM (3.3 Å)
- Summary
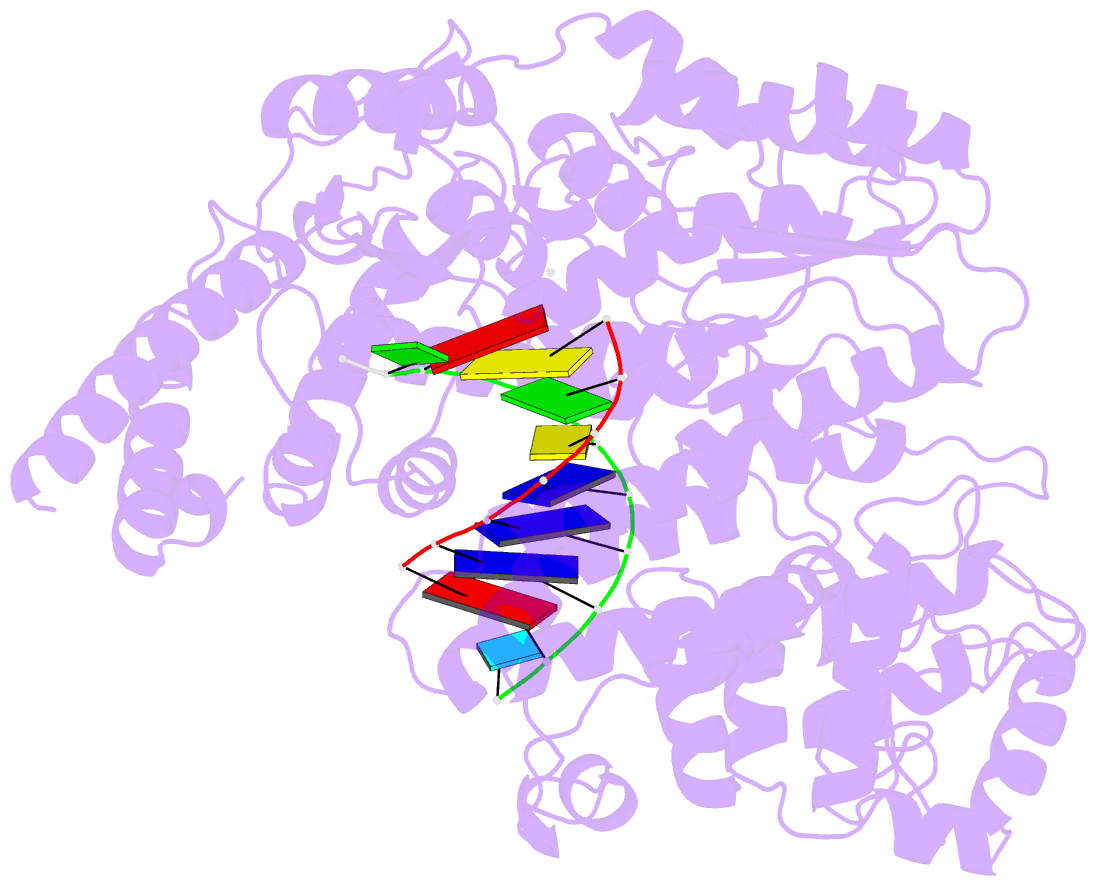
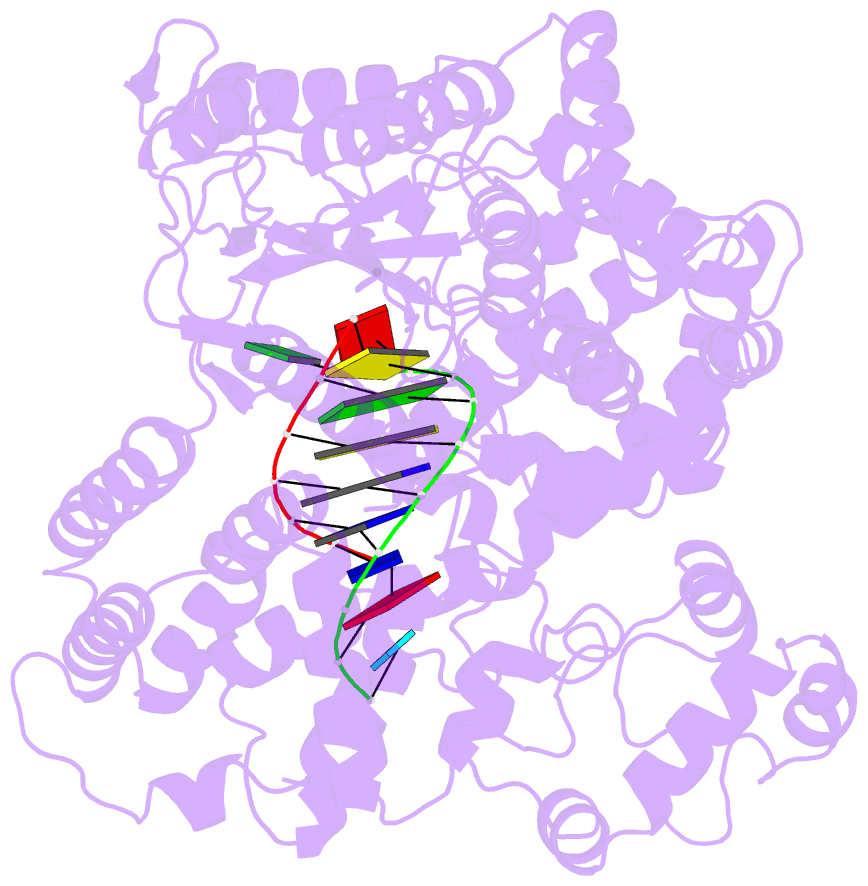
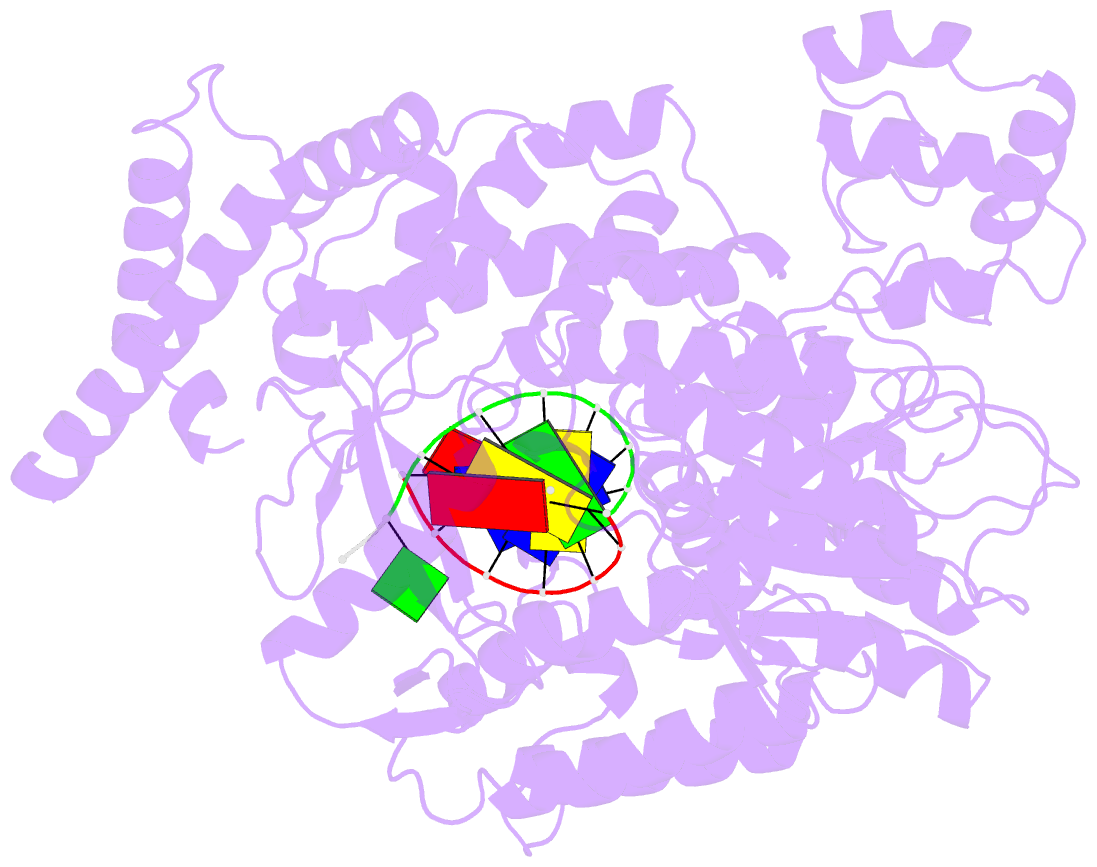
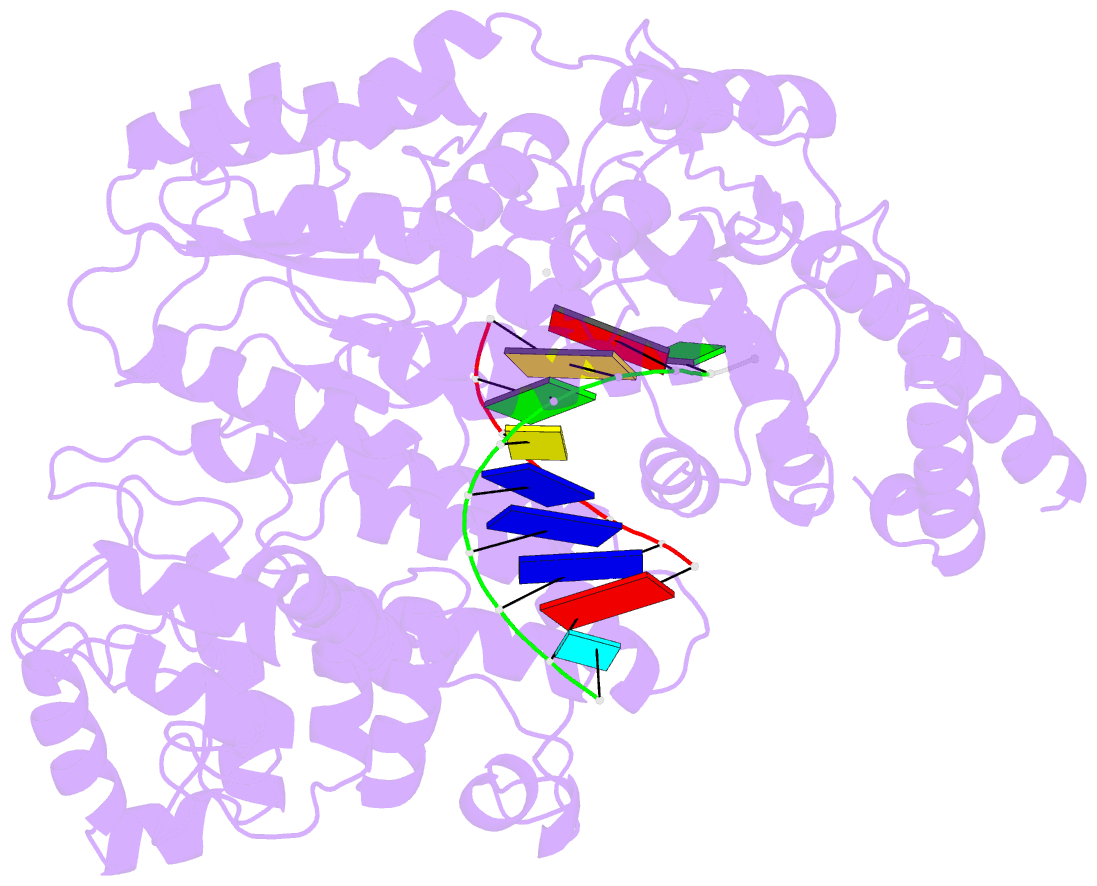
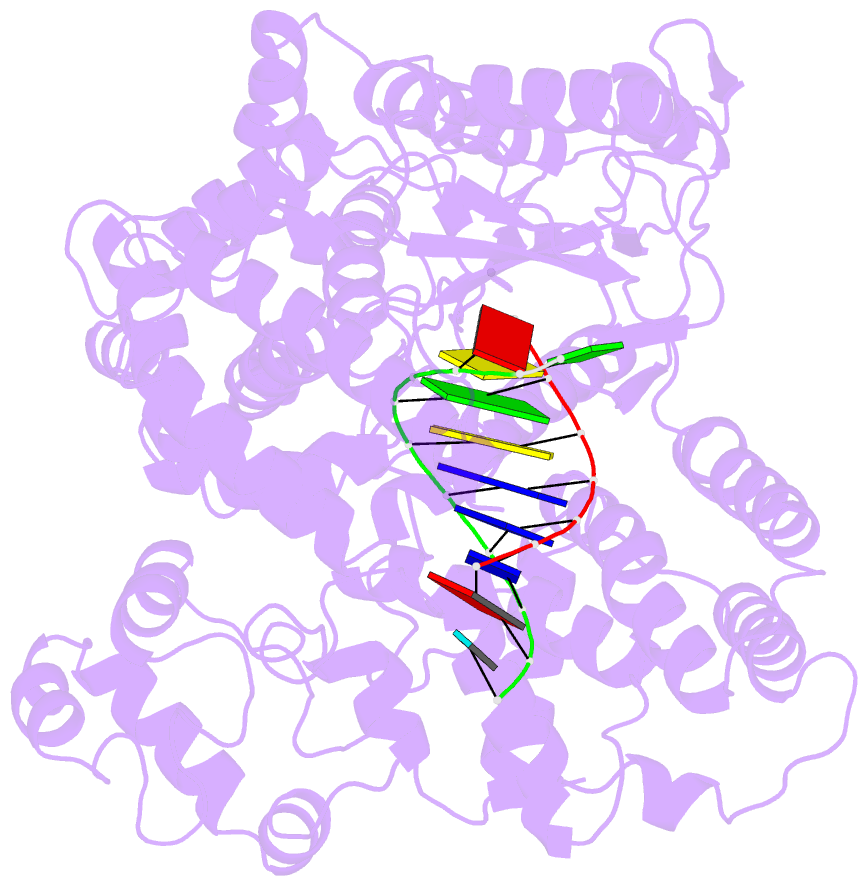
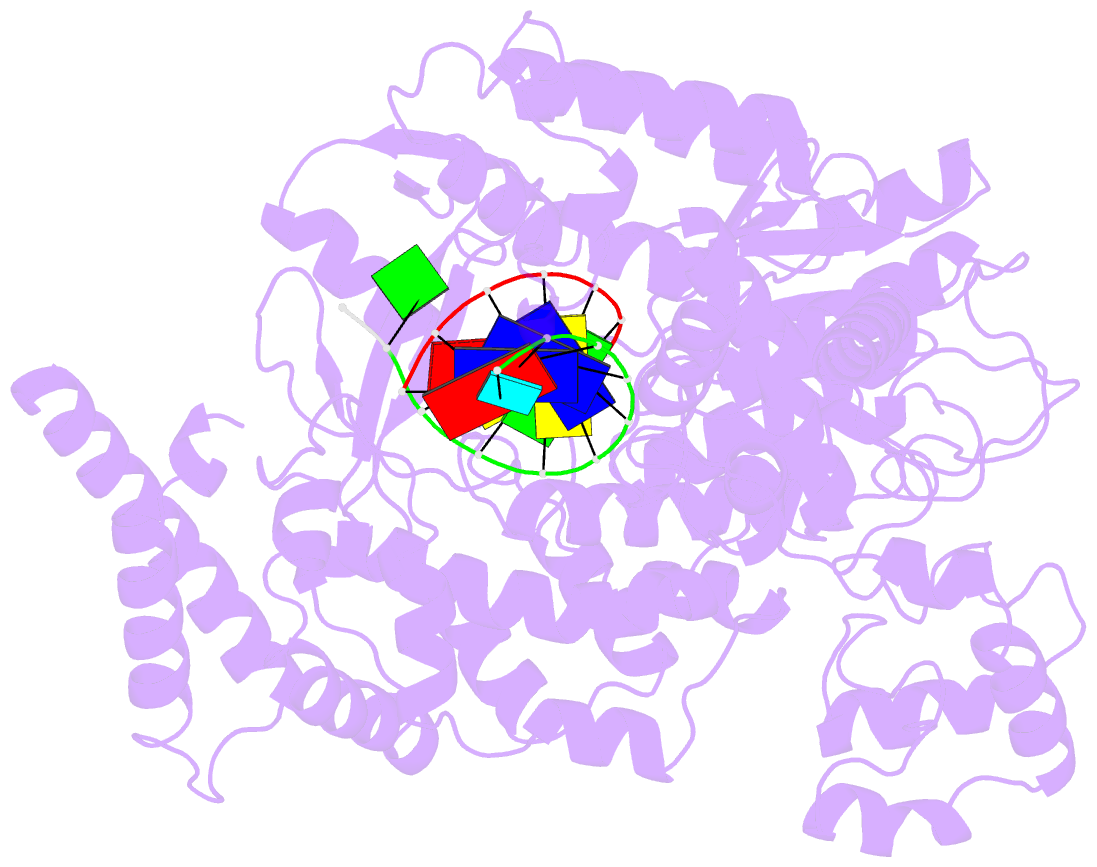
- Structure of line-1 orf2p with template:primer hybrid
- Reference
- Baldwin ET, van Eeuwen T, Hoyos D, Zalevsky A, Tchesnokov EP, Sanchez R, Miller BD, Di Stefano LH, Ruiz FX, Hancock M, Isik E, Mendez-Dorantes C, Walpole T, Nichols C, Wan P, Riento K, Halls-Kass R, Augustin M, Lammens A, Jestel A, Upla P, Xibinaku K, Congreve S, Hennink M, Rogala KB, Schneider AM, Fairman JE, Christensen SM, Desrosiers B, Bisacchi GS, Saunders OL, Hafeez N, Miao W, Kapeller R, Zaller DM, Sali A, Weichenrieder O, Burns KH, Gotte M, Rout MP, Arnold E, Greenbaum BD, Romero DL, LaCava J, Taylor MS (2024): "Structures, functions and adaptations of the human LINE-1 ORF2 protein." Nature, 626, 194-206. doi: 10.1038/s41586-023-06947-z.
- Abstract
- The LINE-1 (L1) retrotransposon is an ancient genetic parasite that has written around one third of the human genome through a "copy-and-paste" mechanism catalyzed by its multifunctional enzyme, open reading frame 2 protein (ORF2p)1. ORF2p reverse transcriptase (RT) and endonuclease activities have been implicated in the pathophysiology of cancer2,3, autoimmunity4,5, and aging6,7, making ORF2p a potential therapeutic target. However, a lack of structural and mechanistic knowledge has hampered efforts to rationally exploit it. We report structures of the human ORF2p 'core' (residues 238-1061, including the RT domain) by X-ray crystallography and cryo-EM in multiple conformational states. Our analyses reveal two novel folded domains, extensive contacts to RNA templates, and associated adaptations that contribute to unique aspects of the L1 replication cycle. Computed integrative structural models of full-length ORF2p show a dynamic closed ring conformation that appears to open during retrotransposition. We characterize ORF2p RT inhibition and reveal its underlying structural basis. Imaging and biochemistry reveal that non-canonical cytosolic ORF2p RT activity can produce RNA:DNA hybrids, activating innate immune signaling via cGAS/STING and resulting in interferon production6-8. In contrast to retroviral RTs, L1 RT is efficiently primed by short RNAs and hairpins, which likely explains cytosolic priming. Additional biochemical activities including processivity, DNA-directed polymerization, non-templated base addition, and template switching together allow us to propose an updated L1 insertion model. Finally, our evolutionary analysis reveals structural conservation between ORF2p and other RNA- and DNA-dependent polymerases. We therefore provide key mechanistic insights into L1 polymerization and insertion, shed light on L1 evolutionary history, and enable rational drug development targeting L1.