Summary information and primary citation
- PDB-id
- 8xc1; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- membrane protein
- Method
- cryo-EM (2.21 Å)
- Summary
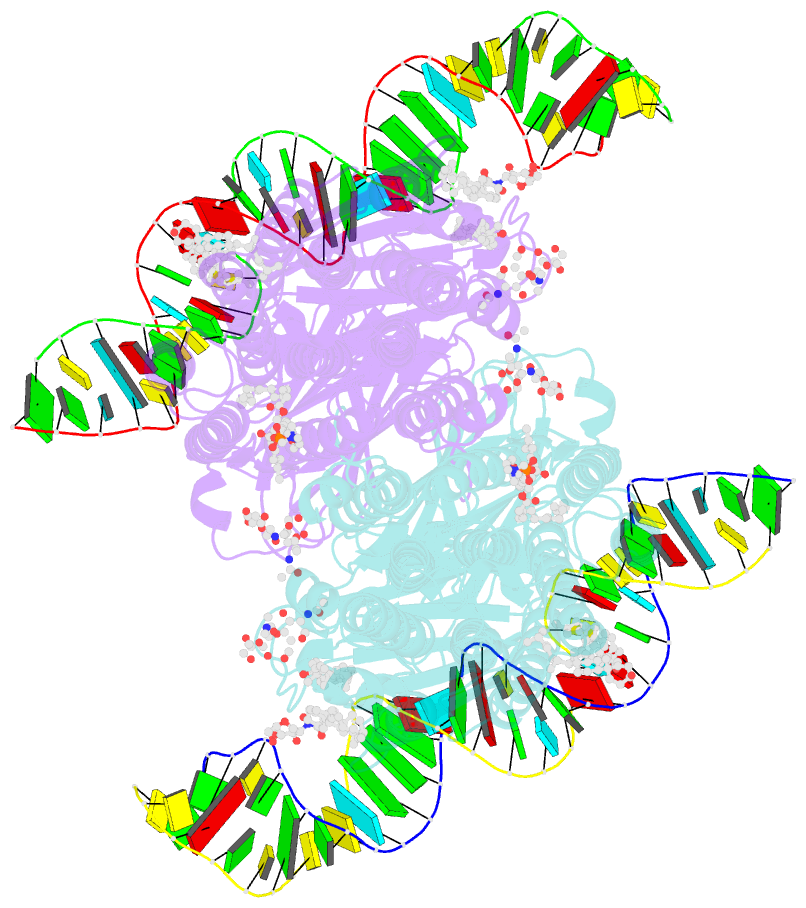
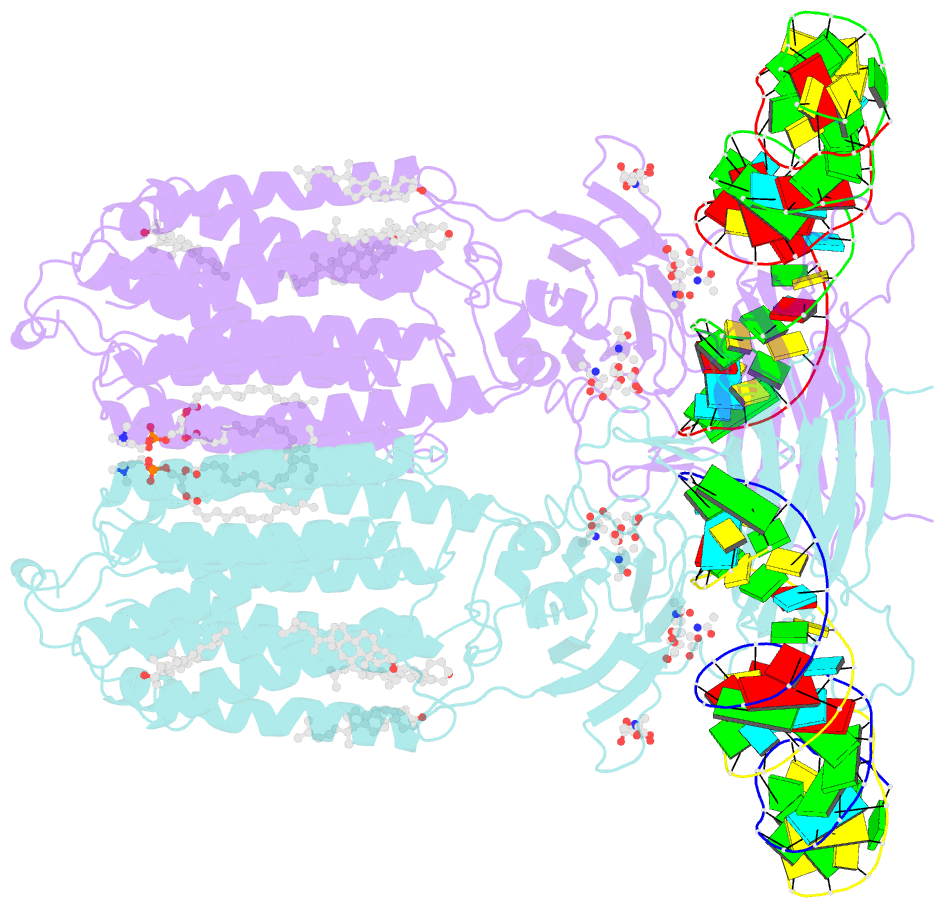
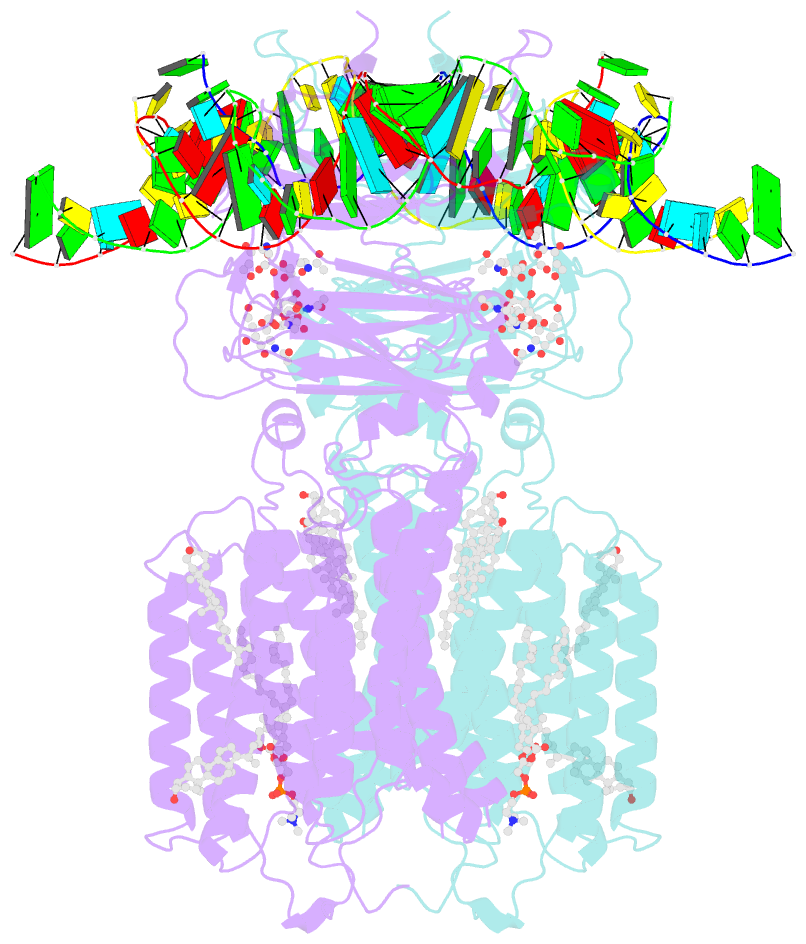
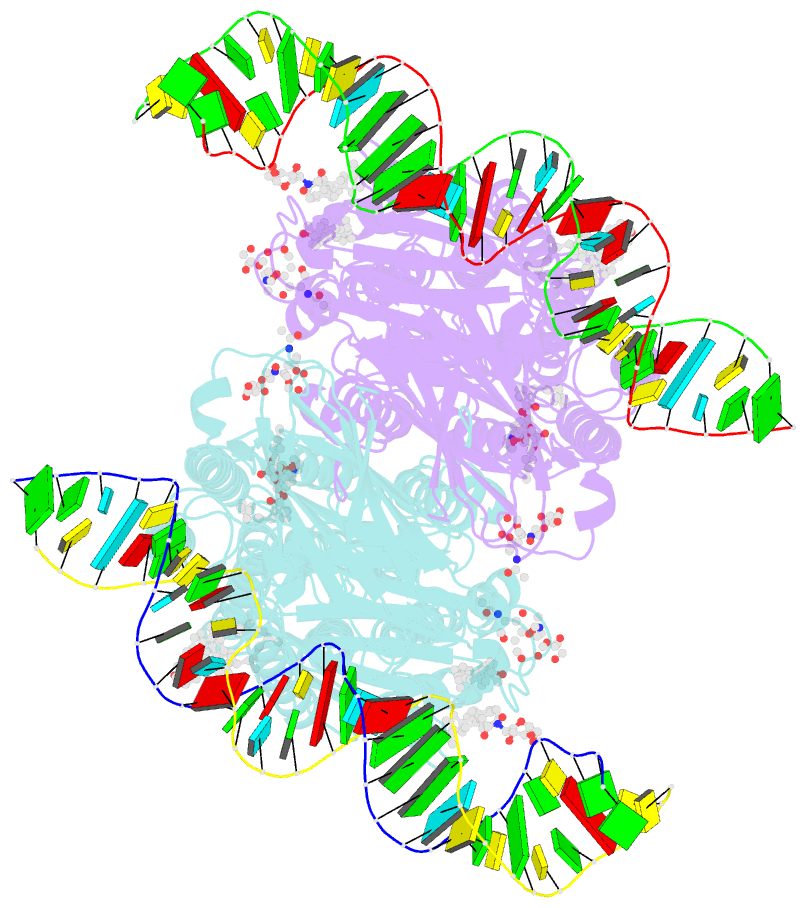
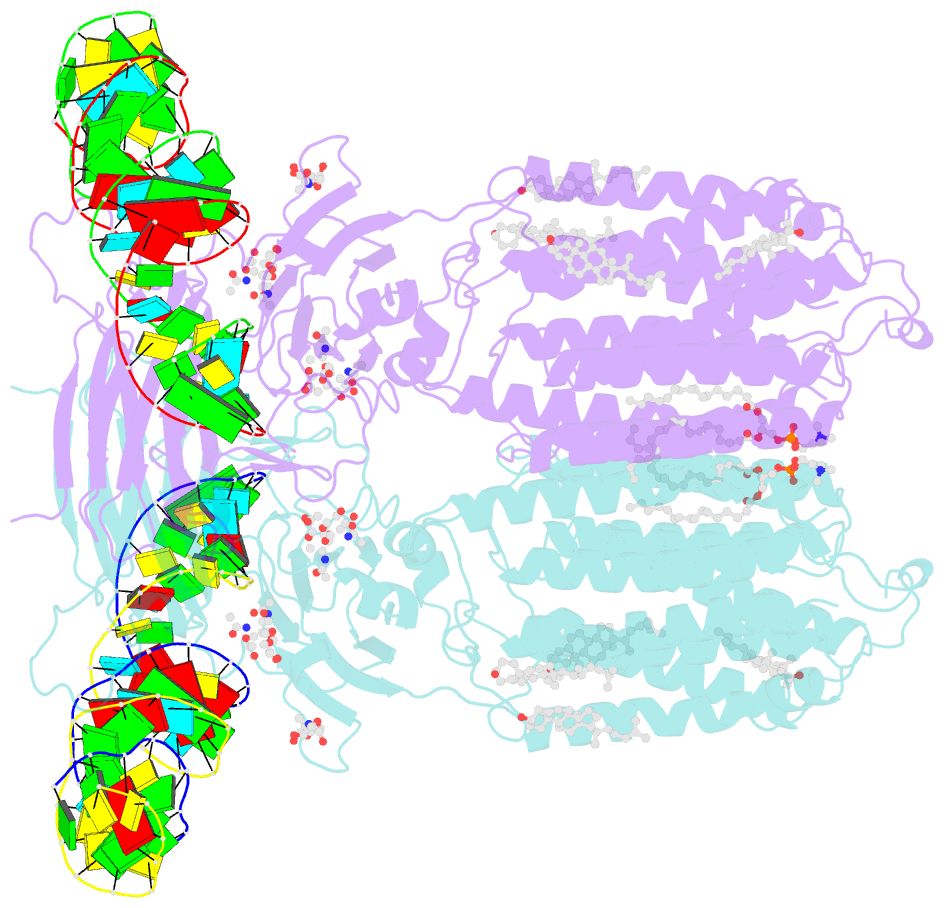
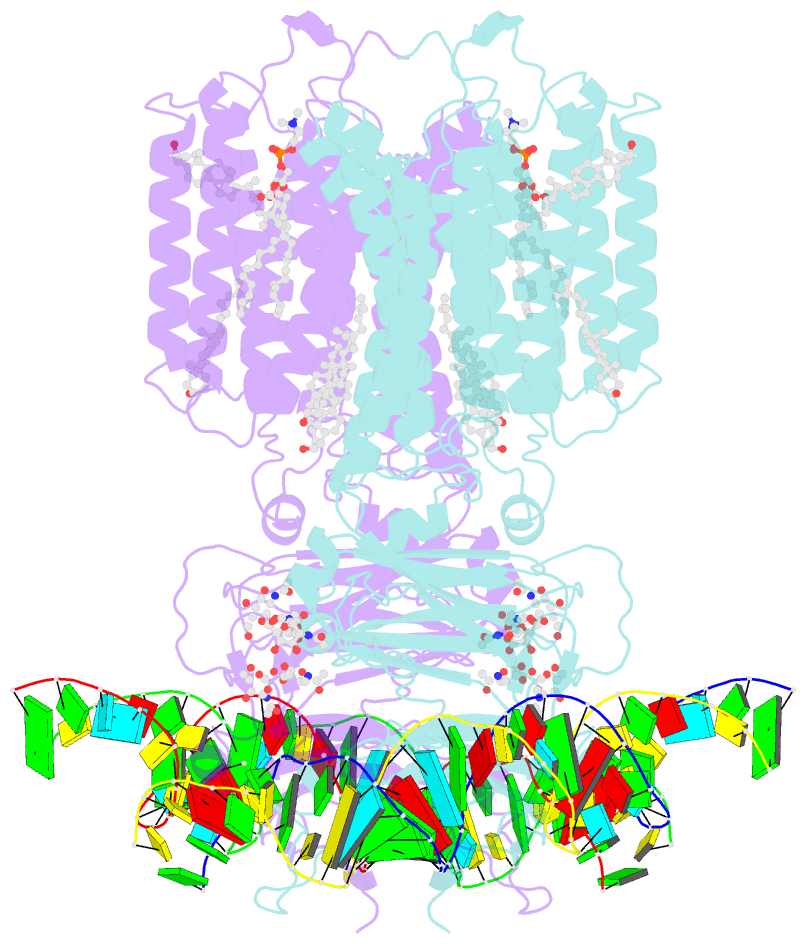
- C. elegans sid1 in complex with dsrna
- Reference
- Wang R, Cong Y, Qian D, Yan C, Gong D (2024): "Structural basis for double-stranded RNA recognition by SID1." Nucleic Acids Res., 52, 6718-6727. doi: 10.1093/nar/gkae395.
- Abstract
- The nucleic acid transport properties of the systemic RNAi-defective (SID) 1 family make them attractive targets for developing RNA-based therapeutics and drugs. However, the molecular basis for double-stranded (ds) RNA recognition by SID1 family remains elusive. Here, we report the cryo-EM structures of Caenorhabditis elegans (c) SID1 alone and in complex with dsRNA, both at a resolution of 2.2 Å. The dimeric cSID1 interacts with two dsRNA molecules simultaneously. The dsRNA is located at the interface between β-strand rich domain (BRD)1 and BRD2 and nearly parallel to the membrane plane. In addition to extensive ionic interactions between basic residues and phosphate backbone, several hydrogen bonds are formed between 2'-hydroxyl group of dsRNA and the contact residues. Additionally, the electrostatic potential surface shows three basic regions are fitted perfectly into three major grooves of dsRNA. These structural characteristics enable cSID1 to bind dsRNA in a sequence-independent manner and to distinguish between DNA and RNA. The cSID1 exhibits no conformational changes upon binding dsRNA, with the exception of a few binding surfaces. Structural mapping of dozens of loss-of-function mutations allows potential interpretation of their diverse functional mechanisms. Our study marks an important step toward mechanistic understanding of the SID1 family-mediated dsRNA uptake.