Summary information and primary citation
- PDB-id
- 8z9h; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- transcription
- Method
- cryo-EM (2.7 Å)
- Summary
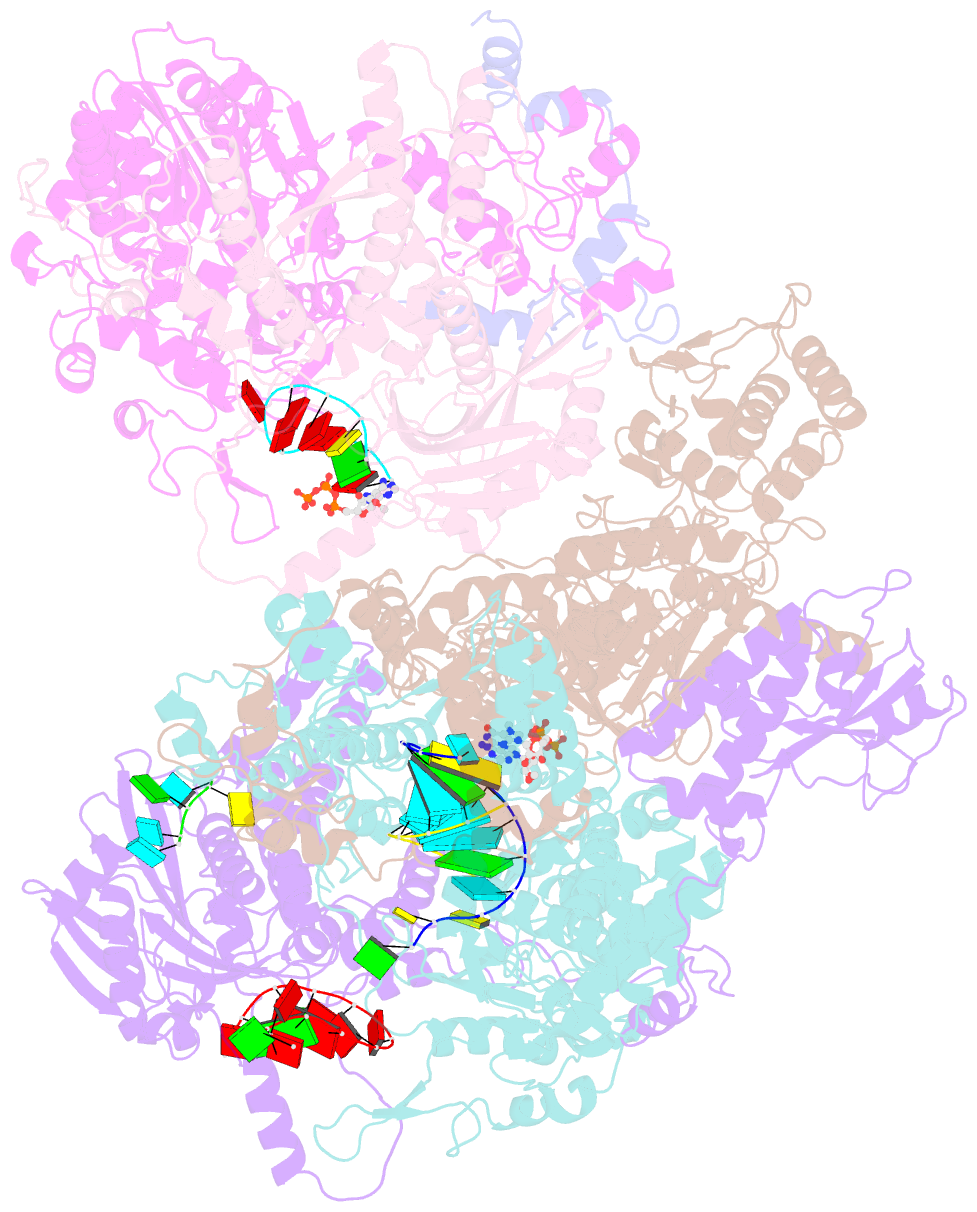
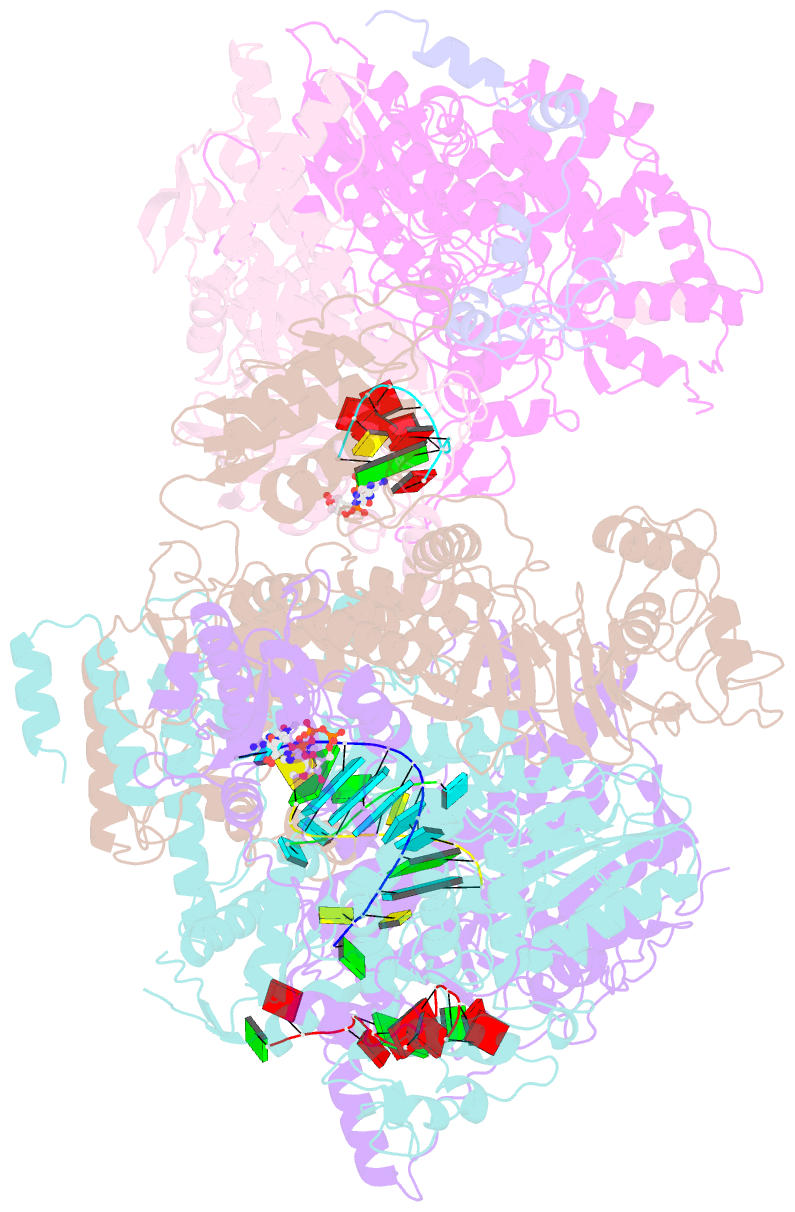
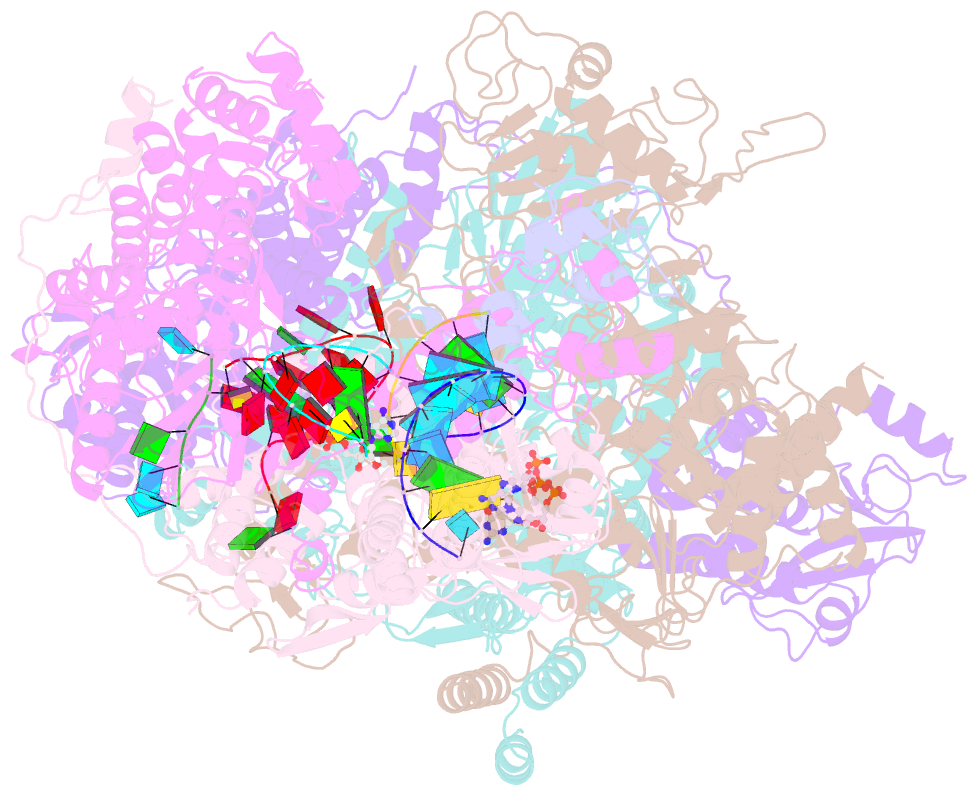
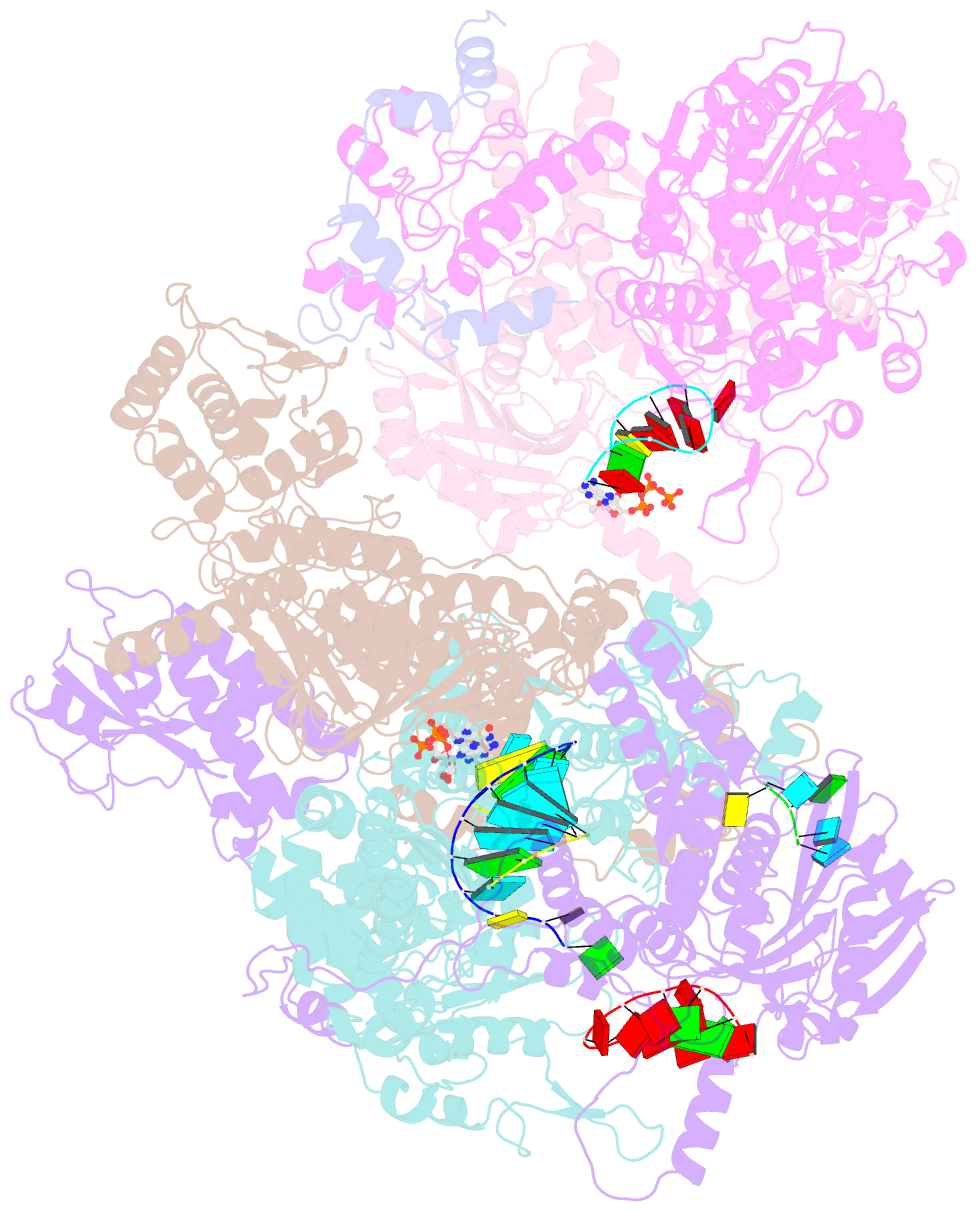
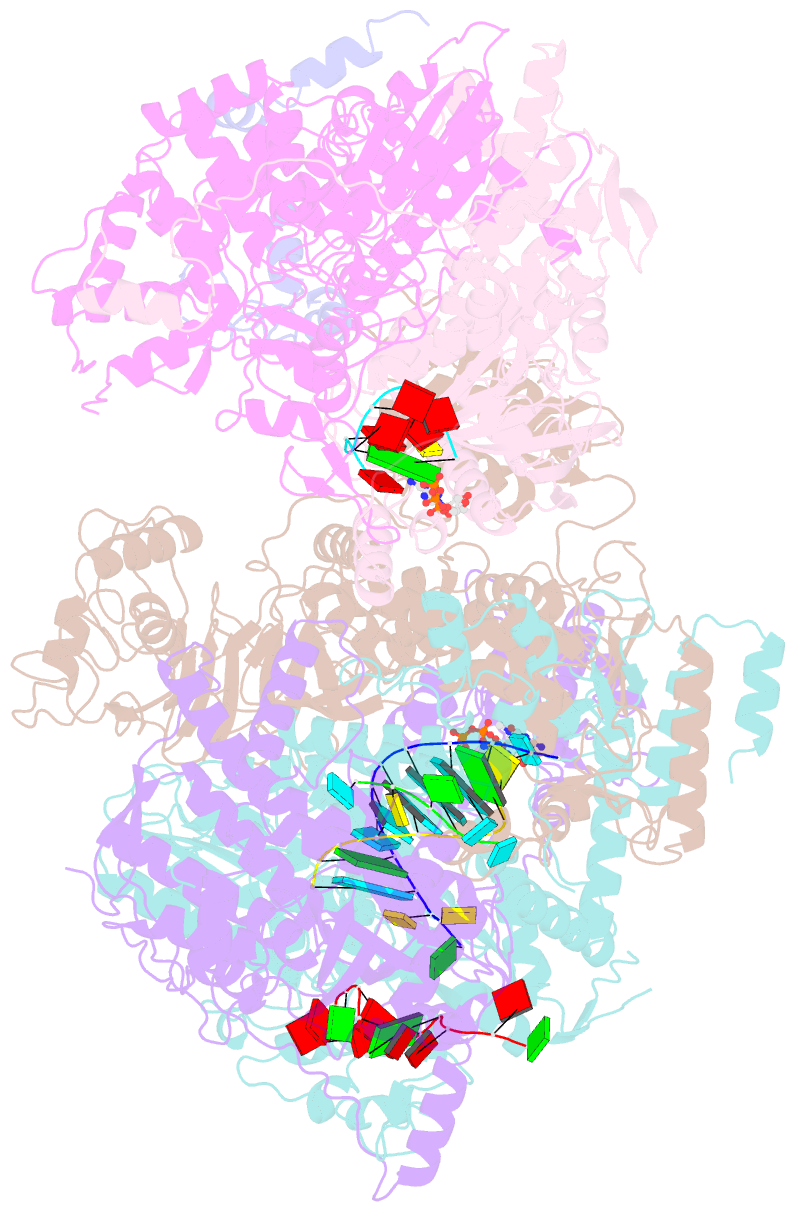
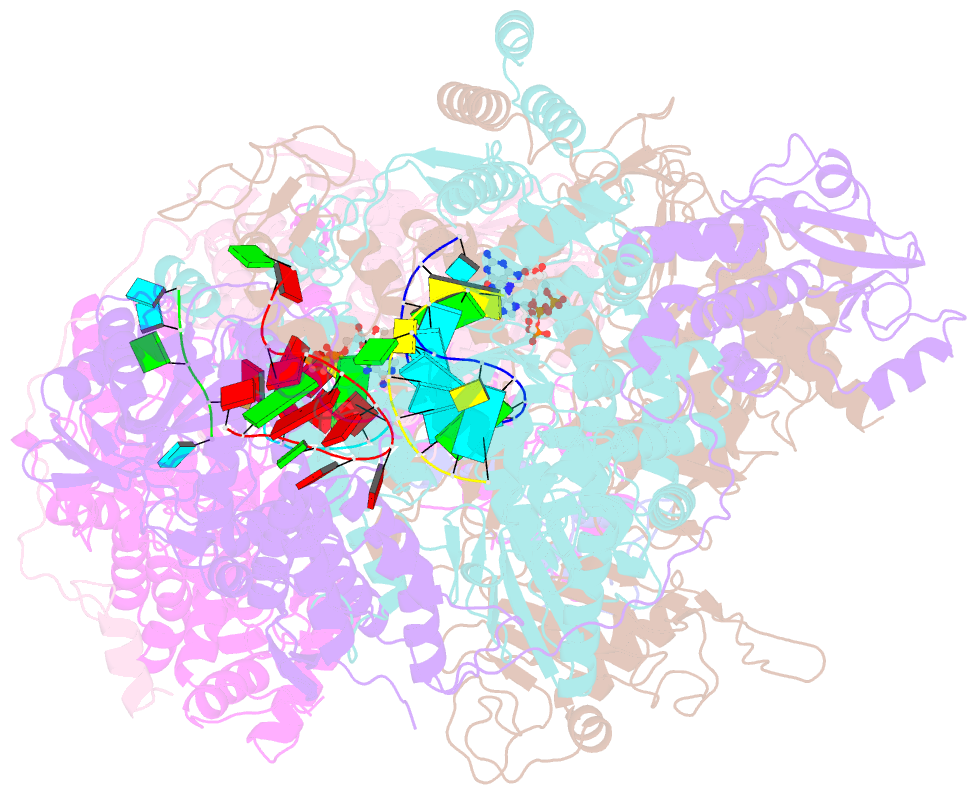
- cryo-EM structure of thogoto virus polymerase in a transcription elongation-reception conformation
- Reference
- Xue L, Chang T, Li Z, Wang C, Zhao H, Li M, Tang P, Wen X, Yu M, Wu J, Bao X, Wang X, Gong P, He J, Chen X, Xiong X (2024): "Cryo-EM structures of Thogoto virus polymerase reveal unique RNA transcription and replication mechanisms among orthomyxoviruses." Nat Commun, 15, 4620. doi: 10.1038/s41467-024-48848-3.
- Abstract
- Influenza viruses and thogotoviruses account for most recognized orthomyxoviruses. Thogotoviruses, exemplified by Thogoto virus (THOV), are capable of infecting humans using ticks as vectors. THOV transcribes mRNA without the extraneous 5' end sequences derived from cap-snatching in influenza virus mRNA. Here, we report cryo-EM structures to characterize THOV polymerase RNA synthesis initiation and elongation. The structures demonstrate that THOV RNA transcription and replication are able to start with short dinucleotide primers and that the polymerase cap-snatching machinery is likely non-functional. Triggered by RNA synthesis, asymmetric THOV polymerase dimers can form without the involvement of host factors. We confirm that, distinctive from influenza viruses, THOV-polymerase RNA synthesis is weakly dependent of the host factors ANP32A/B/E in human cells. This study demonstrates varied mechanisms in RNA synthesis and host factor utilization among orthomyxoviruses, providing insights into the mechanisms behind thogotoviruses' broad-infectivity range.