Summary information and primary citation
- PDB-id
- 9c9m; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- viral protein
- Method
- cryo-EM (2.01 Å)
- Summary
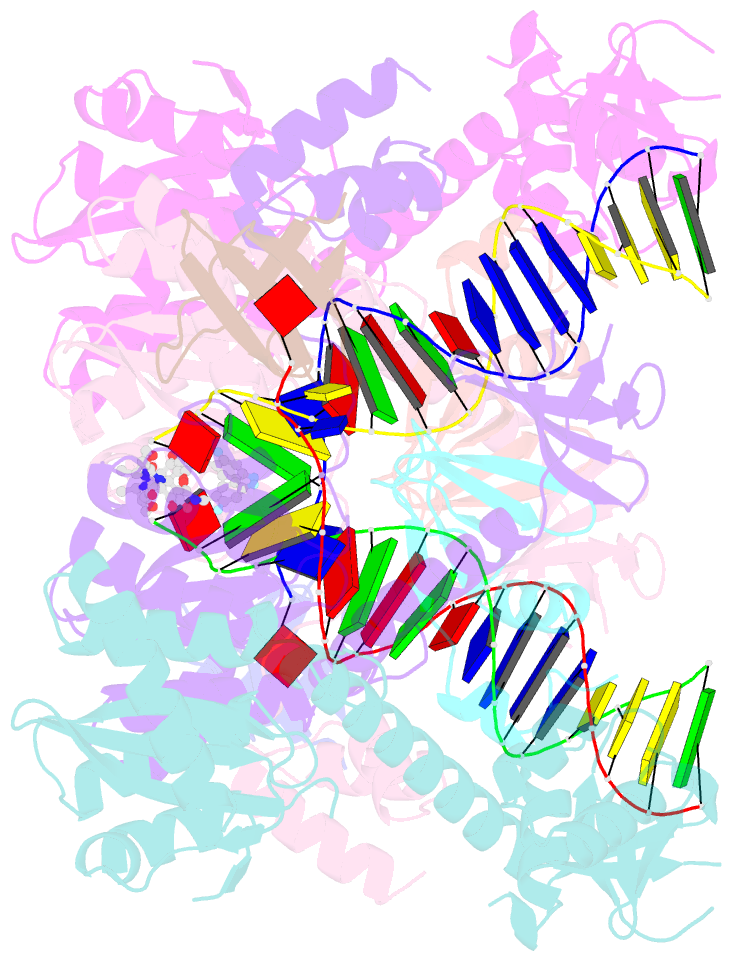
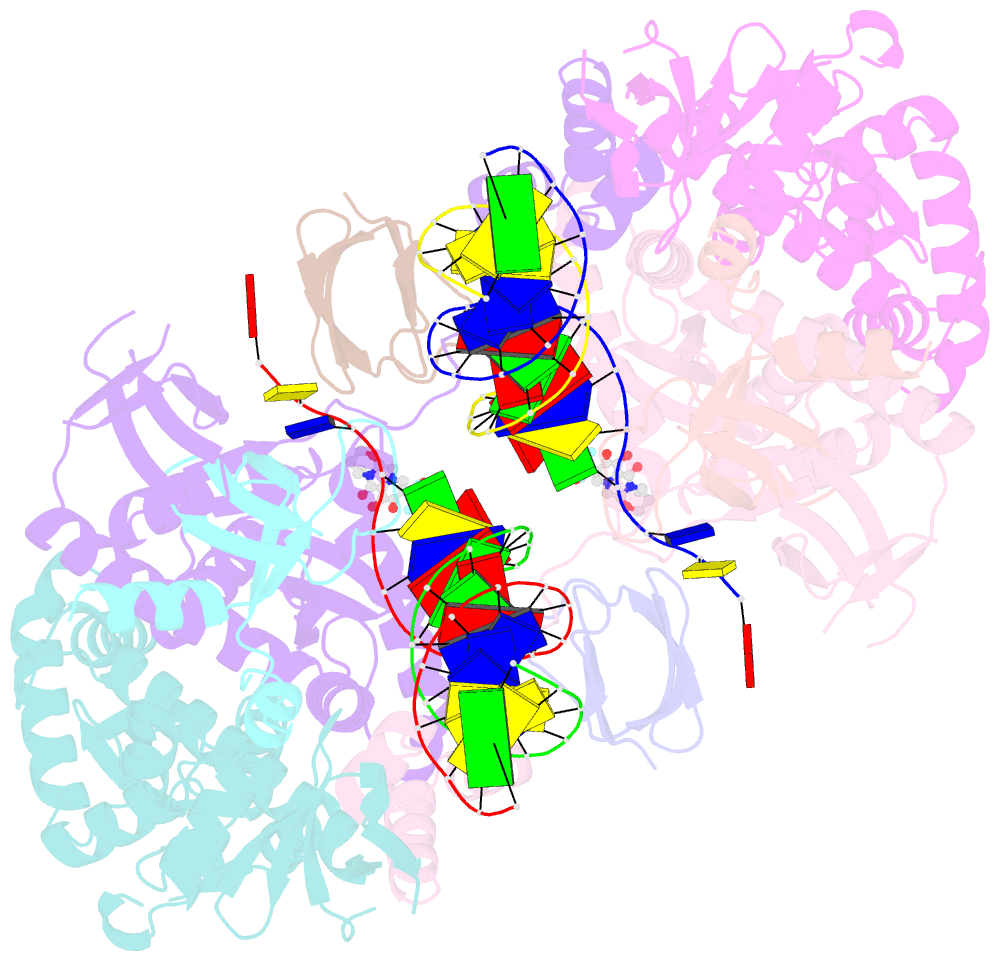
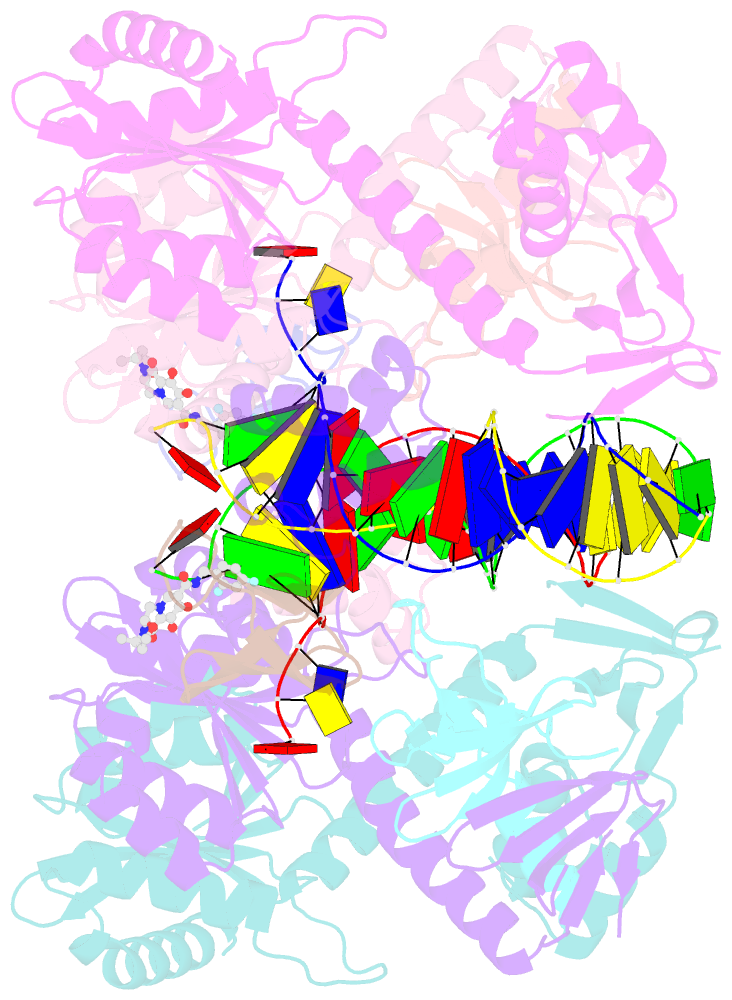
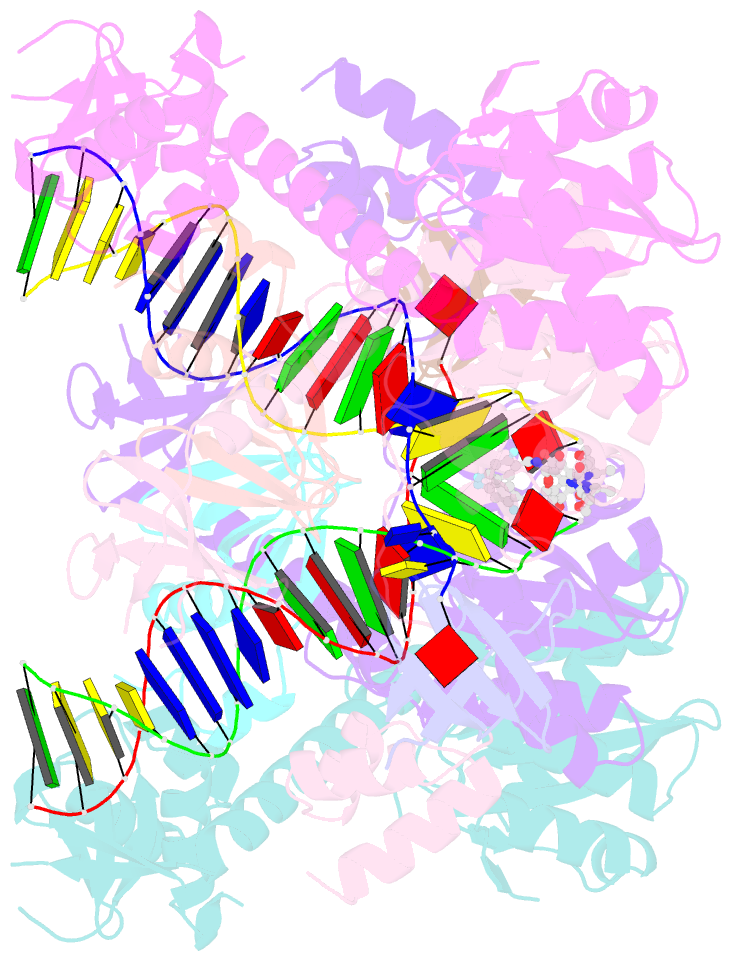
- Hiv-1 intasome core bound with dtg
- Reference
- Li M, Li Z, Chen X, Cui Y, Engelman AN, Craigie R (2024): "HIV-1 Intasomes Assembled with Excess Integrase C-Terminal Domain Protein Facilitate Structural Studies by Cryo-EM and Reveal the Role of the Integrase C-Terminal Tail in HIV-1 Integration." Viruses, 16. doi: 10.3390/v16071166.
- Abstract
- Retroviral integration is mediated by intasome nucleoprotein complexes wherein a pair of viral DNA ends are bridged together by a multimer of integrase (IN). Atomic-resolution structures of HIV-1 intasomes provide detailed insights into the mechanism of integration and inhibition by clinical IN inhibitors. However, previously described HIV-1 intasomes are highly heterogeneous and have the tendency to form stacks, which is a limiting factor in determining high-resolution cryo-EM maps. We have assembled HIV-1 intasomes in the presence of excess IN C-terminal domain protein, which was readily incorporated into the intasomes. The purified intasomes were largely homogeneous and exhibited minimal stacking tendencies. The cryo-EM map resolution was further improved to 2.01 Å, which will greatly facilitate structural studies of IN inhibitor action and drug resistance mechanisms. The C-terminal 18 residues of HIV-1 IN, which are critical for virus replication and integration in vitro, have not been well resolved in previous intasome structures, and its function remains unclear. We show that the C-terminal tail participates in intasome assembly, resides within the intasome core, and forms a small alpha helix (residues 271-276). Mutations that disrupt alpha helix integrity impede IN activity in vitro and disrupt HIV-1 infection at the step of viral DNA integration.